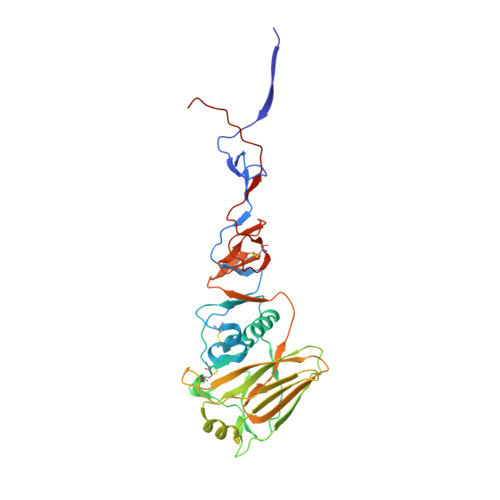
Major antigenic site B of human influenza H3N2 viruses has an evolving local fitness landscape.
Wu, N.C., Otwinowski, J., Thompson, A.J., Nycholat, C.M., Nourmohammad, A., Wilson, I.A.(2020) Nat Commun 11: 1233-1233
- PubMed: 32144244 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-020-15102-5
- Primary Citation Related Structures:
6TZB - PubMed Abstract:
Antigenic drift of influenza virus hemagglutinin (HA) is enabled by facile evolvability. However, HA antigenic site B, which has become immunodominant in recent human H3N2 influenza viruses, is also evolutionarily constrained by its involvement in receptor binding. Here, we employ deep mutational scanning to probe the local fitness landscape of HA antigenic site B in six different human H3N2 strains spanning from 1968 to 2016. We observe that the fitness landscape of HA antigenic site B can be very different between strains. Sequence variants that exhibit high fitness in one strain can be deleterious in another, indicating that the evolutionary constraints of antigenic site B have changed over time. Structural analysis suggests that the local fitness landscape of antigenic site B can be reshaped by natural mutations via modulation of the receptor-binding mode. Overall, these findings elucidate how influenza virus continues to explore new antigenic space despite strong functional constraints.
- Department of Integrative Structural and Computational Biology, The Scripps Research Institute, La Jolla, CA, 92037, USA.
Organizational Affiliation: