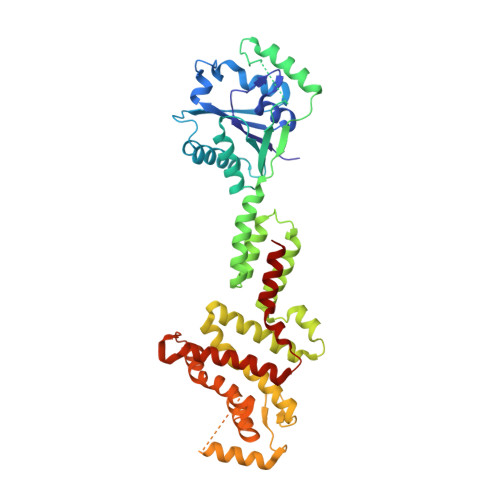
Activation and self-inactivation mechanisms of the cyclic oligoadenylate-dependent CRISPR ribonuclease Csm6.
Garcia-Doval, C., Schwede, F., Berk, C., Rostol, J.T., Niewoehner, O., Tejero, O., Hall, J., Marraffini, L.A., Jinek, M.(2020) Nat Commun 11: 1596-1596
- PubMed: 32221291 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-020-15334-5
- Primary Citation Related Structures:
6TUG - PubMed Abstract:
Bacterial and archaeal CRISPR-Cas systems provide RNA-guided immunity against genetic invaders such as bacteriophages and plasmids. Upon target RNA recognition, type III CRISPR-Cas systems produce cyclic-oligoadenylate second messengers that activate downstream effectors, including Csm6 ribonucleases, via their CARF domains. Here, we show that Enteroccocus italicus Csm6 (EiCsm6) degrades its cognate cyclic hexa-AMP (cA6) activator, and report the crystal structure of EiCsm6 bound to a cA6 mimic. Our structural, biochemical, and in vivo functional assays reveal how cA6 recognition by the CARF domain activates the Csm6 HEPN domains for collateral RNA degradation, and how CARF domain-mediated cA6 cleavage provides an intrinsic off-switch to limit Csm6 activity in the absence of ring nucleases. These mechanisms facilitate rapid invader clearance and ensure termination of CRISPR interference to limit self-toxicity.
- Department of Biochemistry, University of Zurich, Winterthurerstrasse 190, CH-8057, Zurich, Switzerland.
Organizational Affiliation: