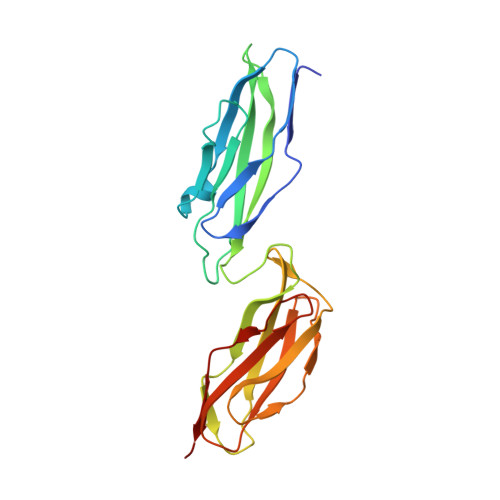
The extracellular juncture domains in the intimin passenger adopt a constitutively extended conformation inducing restraints to its sphere of action.
Weikum, J., Kulakova, A., Tesei, G., Yoshimoto, S., Jaegerum, L.V., Schutz, M., Hori, K., Skepo, M., Harris, P., Leo, J.C., Morth, J.P.(2020) Sci Rep 10: 21249-21249
- PubMed: 33277518 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-020-77706-7
- Primary Citation Related Structures:
6TPL, 6TQD - PubMed Abstract:
Enterohemorrhagic and enteropathogenic Escherichia coli are among the most important food-borne pathogens, posing a global health threat. The virulence factor intimin is essential for the attachment of pathogenic E. coli to the intestinal host cell. Intimin consists of four extracellular bacterial immunoglobulin-like (Big) domains, D00-D2, extending into the fifth lectin subdomain (D3) that binds to the Tir-receptor on the host cell. Here, we present the crystal structures of the elusive D00-D0 domains at 1.5 Å and D0-D1 at 1.8 Å resolution, which confirms that the passenger of intimin has five distinct domains. We describe that D00-D0 exhibits a higher degree of rigidity and D00 likely functions as a juncture domain at the outer membrane-extracellular medium interface. We conclude that D00 is a unique Big domain with a specific topology likely found in a broad range of other inverse autotransporters. The accumulated data allows us to model the complete passenger of intimin and propose functionality to the Big domains, D00-D0-D1, extending directly from the membrane.
- Membrane Transport Group, Centre for Molecular Medicine Norway (NCMM), Nordic EMBL Partnership, University of Oslo, P.O. Box 1137, Blindern, 0318, Oslo, Norway.
Organizational Affiliation: