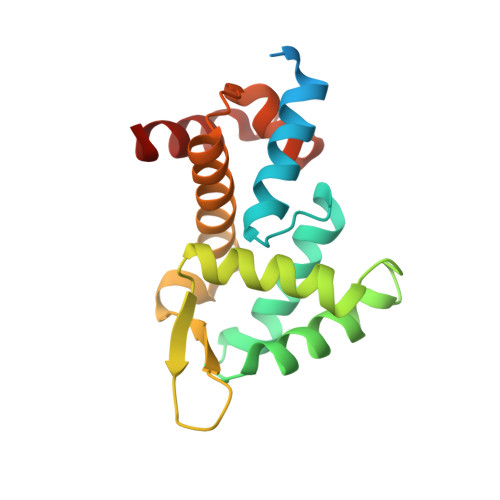
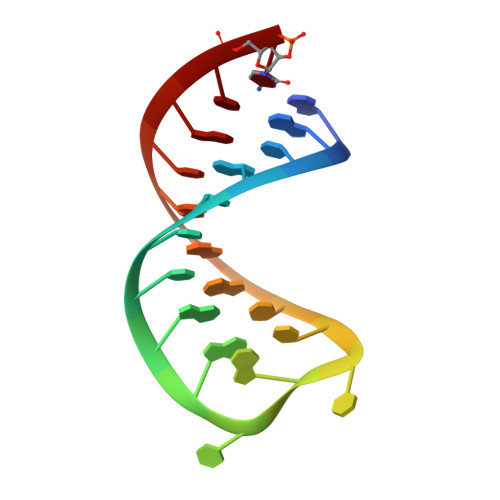
Structural basis for the recognition of transiently structured AU-rich elements by Roquin.
Binas, O., Tants, J.N., Peter, S.A., Janowski, R., Davydova, E., Braun, J., Niessing, D., Schwalbe, H., Weigand, J.E., Schlundt, A.(2020) Nucleic Acids Res 48: 7385-7403
- PubMed: 32491174 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkaa465
- Primary Citation Related Structures:
6TQA, 6TQB, 6XWJ, 6XWW, 6XXA, 6XXB - PubMed Abstract:
Adenylate/uridylate-rich elements (AREs) are the most common cis-regulatory elements in the 3'-untranslated region (UTR) of mRNAs, where they fine-tune turnover by mediating mRNA decay. They increase plasticity and efficacy of mRNA regulation and are recognized by several ARE-specific RNA-binding proteins (RBPs). Typically, AREs are short linear motifs with a high content of complementary A and U nucleotides and often occur in multiple copies. Although thermodynamically rather unstable, the high AU-content might enable transient secondary structure formation and modify mRNA regulation by RBPs. We have recently suggested that the immunoregulatory RBP Roquin recognizes folded AREs as constitutive decay elements (CDEs), resulting in shape-specific ARE-mediated mRNA degradation. However, the structural evidence for a CDE-like recognition of AREs by Roquin is still lacking. We here present structures of CDE-like folded AREs, both in their free and protein-bound form. Moreover, the AREs in the UCP3 3'-UTR are additionally bound by the canonical ARE-binding protein AUF1 in their linear form, adopting an alternative binding-interface compared to the recognition of their CDE structure by Roquin. Strikingly, our findings thus suggest that AREs can be recognized in multiple ways, allowing control over mRNA regulation by adapting distinct conformational states, thus providing differential accessibility to regulatory RBPs.
- Institute for Organic Chemistry and Chemical Biology, Goethe University Frankfurt and Center for Biomolecular Magnetic Resonance (BMRZ), 60438 Frankfurt, Germany.
Organizational Affiliation: