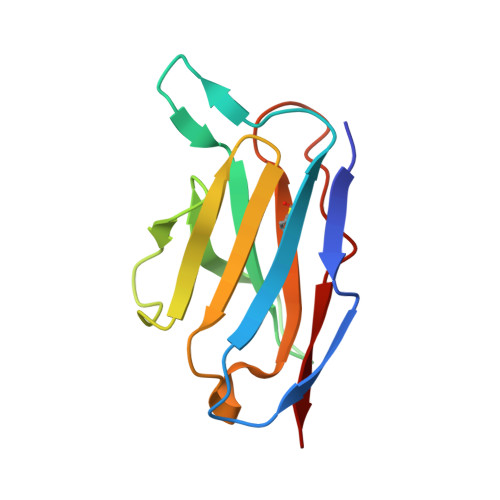
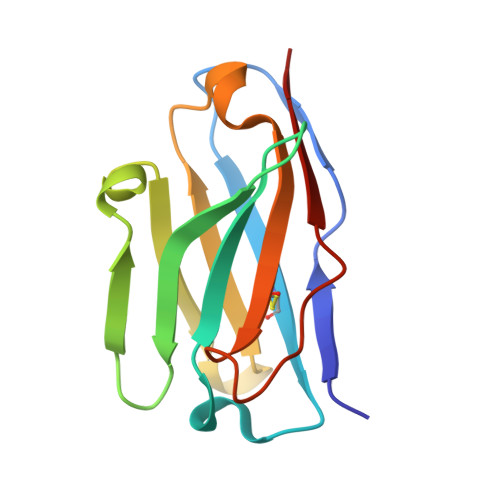
Structural characterization of an unprecedented lectin-like antitumoral anti-MUC1 antibody.
Macias-Leon, J., Bermejo, I.A., Asin, A., Garcia-Garcia, A., Companon, I., Jimenez-Moreno, E., Coelho, H., Mangini, V., Albuquerque, I.S., Marcelo, F., Asensio, J.L., Bernardes, G.J.L., Joshi, H.J., Fiammengo, R., Blixt, O., Hurtado-Guerrero, R., Corzana, F.(2020) Chem Commun (Camb) 56: 15137-15140
- PubMed: 33211039 Search on PubMed
- DOI: https://doi.org/10.1039/d0cc06349e
- Primary Citation Related Structures:
6TNP - PubMed Abstract:
The molecular basis of antibody 5E5, which recognizes the entire GalNAc unit as a primary epitope is disclosed. The antibody's contacts with the peptide are mostly limited to two residues, allowing it to show some degree of promiscuity. These findings open the door to the chemical design of peptide-mimetics for developing efficient anti-cancer vaccines and diagnostic tools.
- Institute of Biocomputation and Physics of Complex Systems (BIFI), University of Zaragoza, Mariano Esquillor s/n, Campus Rio Ebro, Edificio I+D, Zaragoza, Spain. rhurtado@bifi.es.
Organizational Affiliation: