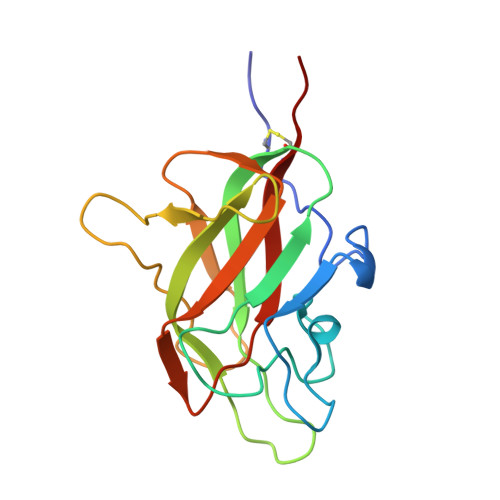
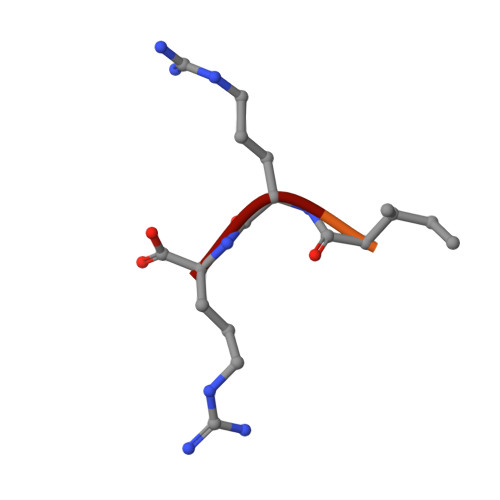
VEGFA, B, C: Implications of the C-Terminal Sequence Variations for the Interaction with Neuropilins.
Eldrid, C., Zloh, M., Fotinou, C., Yelland, T., Yu, L., Mota, F., Selwood, D.L., Djordjevic, S.(2022) Biomolecules 12
- PubMed: 35327564 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/biom12030372
- Primary Citation Related Structures:
6TDB, 6TJT, 6TKK - PubMed Abstract:
Vascular endothelial growth factors (VEGFs) are the key regulators of blood and lymphatic vessels' formation and function. Each of the proteins from the homologous family VEGFA, VEGFB, VEGFC and VEGFD employs a core cysteine-knot structural domain for the specific interaction with one or more of the cognate tyrosine kinase receptors. Additional diversity is exhibited by the involvement of neuropilins-transmembrane co-receptors, whose b1 domain contains the binding site for the C-terminal sequence of VEGFs. Although all relevant isoforms of VEGFs that interact with neuropilins contain the required C-terminal Arg residue, there is selectivity of neuropilins and VEGF receptors for the VEGF proteins, which is reflected in the physiological roles that they mediate. To decipher the contribution made by the C-terminal sequences of the individual VEGF proteins to that functional differentiation, we determined structures of molecular complexes of neuropilins and VEGF-derived peptides and examined binding interactions for all neuropilin-VEGF pairs experimentally and computationally. While X-ray crystal structures and ligand-binding experiments highlighted similarities between the ligands, the molecular dynamics simulations uncovered conformational preferences of VEGF-derived peptides beyond the C-terminal arginine that contribute to the ligand selectivity of neuropilins. The implications for the design of the selective antagonists of neuropilins' functions are discussed.
- Structural and Molecular Biology, ISMB, Division of Biosciences, University College London, Gower Street, London WC1E 6BT, UK.
Organizational Affiliation: