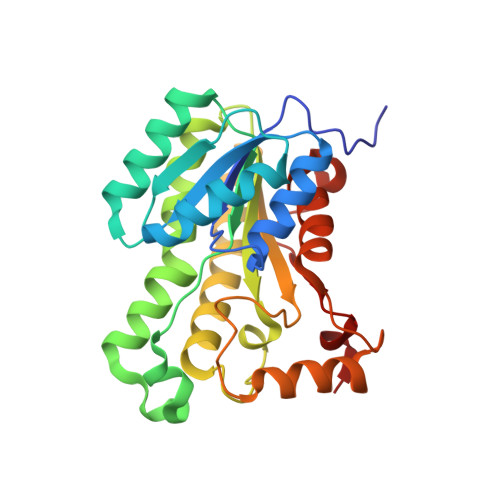
The Kalimantacin Polyketide Antibiotics Inhibit Fatty Acid Biosynthesis in Staphylococcus aureus by Targeting the Enoyl-Acyl Carrier Protein Binding Site of FabI.
Fage, C.D., Lathouwers, T., Vanmeert, M., Gao, L.J., Vrancken, K., Lammens, E.M., Weir, A.N.M., Degroote, R., Cuppens, H., Kosol, S., Simpson, T.J., Crump, M.P., Willis, C.L., Herdewijn, P., Lescrinier, E., Lavigne, R., Anne, J., Masschelein, J.(2020) Angew Chem Int Ed Engl 59: 10549-10556
- PubMed: 32208550 Search on PubMed
- DOI: https://doi.org/10.1002/anie.201915407
- Primary Citation Related Structures:
6TBB, 6TBC - PubMed Abstract:
The enoyl-acyl carrier protein reductase enzyme FabI is essential for fatty acid biosynthesis in Staphylococcus aureus and represents a promising target for the development of novel, urgently needed anti-staphylococcal agents. Here, we elucidate the mode of action of the kalimantacin antibiotics, a novel class of FabI inhibitors with clinically-relevant activity against multidrug-resistant S. aureus. By combining X-ray crystallography with molecular dynamics simulations, in vitro kinetic studies and chemical derivatization experiments, we characterize the interaction between the antibiotics and their target, and we demonstrate that the kalimantacins bind in a unique conformation that differs significantly from the binding mode of other known FabI inhibitors. We also investigate mechanisms of acquired resistance in S. aureus and identify key residues in FabI that stabilize the binding of the antibiotics. Our findings provide intriguing insights into the mode of action of a novel class of FabI inhibitors that will inspire future anti-staphylococcal drug development.
- Department of Chemistry, University of Warwick, Coventry, CV4 7AL, UK.
Organizational Affiliation: