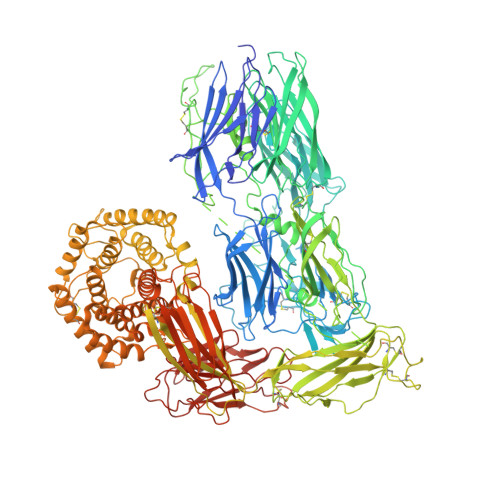
Cryo-EM structures show the mechanistic basis of pan-peptidase inhibition by human alpha 2 -macroglobulin.
Luque, D., Goulas, T., Mata, C.P., Mendes, S.R., Gomis-Ruth, F.X., Caston, J.R.(2022) Proc Natl Acad Sci U S A 119: e2200102119-e2200102119
- PubMed: 35500114 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2200102119
- Primary Citation Related Structures:
6TAV, 7O7L, 7O7M, 7O7N, 7O7O, 7O7P, 7O7Q, 7O7R, 7O7S - PubMed Abstract:
Human α2-macroglobulin (hα2M) is a multidomain protein with a plethora of essential functions, including transport of signaling molecules and endopeptidase inhibition in innate immunity. Here, we dissected the molecular mechanism of the inhibitory function of the ∼720-kDa hα2M tetramer through eight cryo–electron microscopy (cryo-EM) structures of complexes from human plasma. In the native complex, the hα2M subunits are organized in two flexible modules in expanded conformation, which enclose a highly porous cavity in which the proteolytic activity of circulating plasma proteins is tested. Cleavage of bait regions exposed inside the cavity triggers rearrangement to a compact conformation, which closes openings and entraps the prey proteinase. After the expanded-to-compact transition, which occurs independently in the four subunits, the reactive thioester bond triggers covalent linking of the proteinase, and the receptor-binding domain is exposed on the tetramer surface for receptor-mediated clearance from circulation. These results depict the molecular mechanism of a unique suicidal inhibitory trap.
- Spanish National Microbiology Centre, Institute of Health Carlos III, 28220 Madrid, Spain.
Organizational Affiliation: