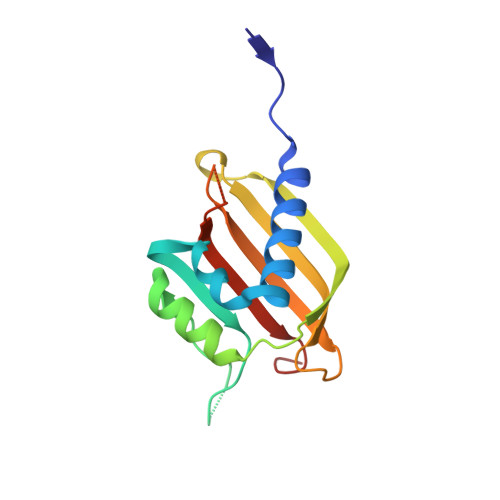
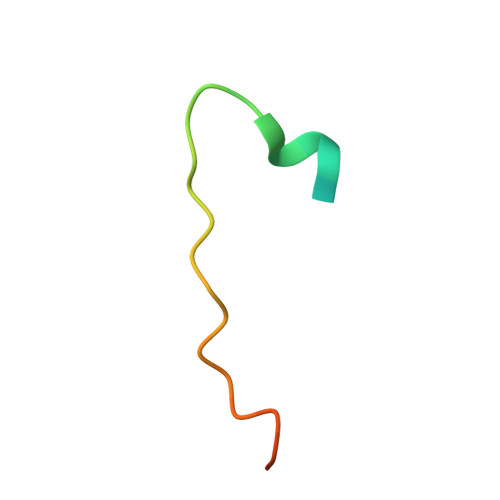
Caprin-1 binding to the critical stress granule protein G3BP1 is regulated by pH
Schulte, T., Panas, M.D., Williams, L., Kedersha, N., Fleck, J.S., Tan, T.J., Olsson, A., Morro, A.M., Hanke, L., Nilvebrant, J., Giang, K.A., Nygren, P.A., Anderson, P., Achour, A., McInerney, G.M.(2021) bioRxiv