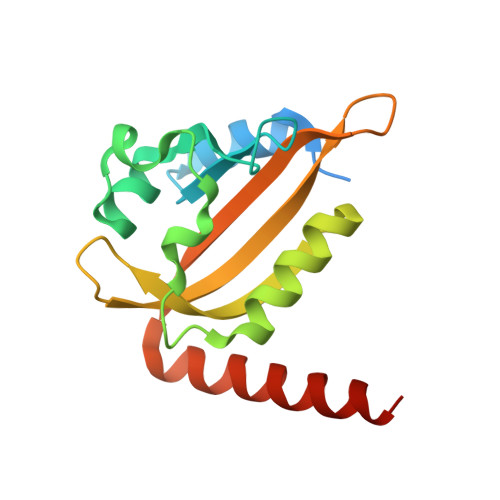
An Optogenetic Tool for Induced Protein Stabilization Based on the Phaeodactylum tricornutum Aureochrome 1a Light-Oxygen-Voltage Domain.
Hepp, S., Trauth, J., Hasenjager, S., Bezold, F., Essen, L.O., Taxis, C.(2020) J Mol Biology 432: 1880-1900
- PubMed: 32105734 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2020.02.019
- Primary Citation Related Structures:
6T73, 6T74 - PubMed Abstract:
Control of cellular events by optogenetic tools is a powerful approach to manipulate cellular functions in a minimally invasive manner. A common problem posed by the application of optogenetic tools is to tune the activity range to be physiologically relevant. Here, we characterized a photoreceptor of the light-oxygen-voltage (LOV) domain family of Phaeodactylum tricornutum aureochrome 1a (AuLOV) as a tool for increasing protein stability under blue light conditions in budding yeast. Structural studies of AuLOV wt , the variants AuLOV M254 , and AuLOV W349 revealed alternative dimer association modes for the dark state, which differ from previously reported AuLOV dark-state structures. Rational design of AuLOV-dimer interface mutations resulted in an optimized optogenetic tool that we fused to the photoactivatable adenylyl cyclase from Beggiatoa sp. This synergistic light-regulation approach using two photoreceptors resulted in an optimized, photoactivatable adenylyl cyclase with a cyclic adenosine monophosphate production activity that matches the physiological range of Saccharomyces cerevisiae. Overall, we enlarged the optogenetic toolbox for yeast and demonstrated the importance of fine-tuning the optogenetic tool activity for successful application in cells.
- Unit for Structural Biochemistry, Department of Chemistry, Philipps-Universität Marburg, Hans-Meerwein-Strasse 4, 35032 Marburg, Germany; Center of Synthetic Microbiology, Philipps Universität Marburg, Hans-Meerwein- Strasse 4, 35032 Marburg, Germany.
Organizational Affiliation: