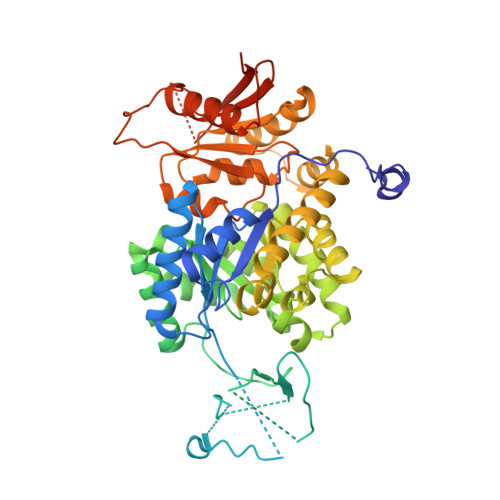
Structural and kinetic characterization of Trypanosoma congolense pyruvate kinase.
Pinto Torres, J.E., Yuan, M., Goossens, J., Versees, W., Caljon, G., Michels, P.A., Walkinshaw, M.D., Magez, S., Sterckx, Y.G.(2020) Mol Biochem Parasitol 236: 111263-111263
- PubMed: 32084384 Search on PubMed
- DOI: https://doi.org/10.1016/j.molbiopara.2020.111263
- Primary Citation Related Structures:
6SU1, 6SU2 - PubMed Abstract:
Trypanosoma are blood-borne parasites and are the causative agents of neglected tropical diseases (NTDs) affecting both humans and animals. These parasites mainly rely on glycolysis for their energy production within the mammalian host, which is why trypanosomal glycolytic enzymes have been pursued as interesting targets for the development of trypanocidal drugs. The structure-function relationships of pyruvate kinases (PYKs) from trypanosomatids (Trypanosoma and Leishmania) have been well-studied within this context. In this paper, we describe the structural and enzymatic characterization of PYK from T. congolense (TcoPYK), the main causative agent of Animal African Trypanosomosis (AAT), by employing a combination of enzymatic assays, thermal unfolding studies and X-ray crystallography.
- Research Unit for Cellular and Molecular Immunology (CMIM), Vrije Universiteit Brussel, Pleinlaan 2, B-1050 Brussels, Belgium.
Organizational Affiliation: