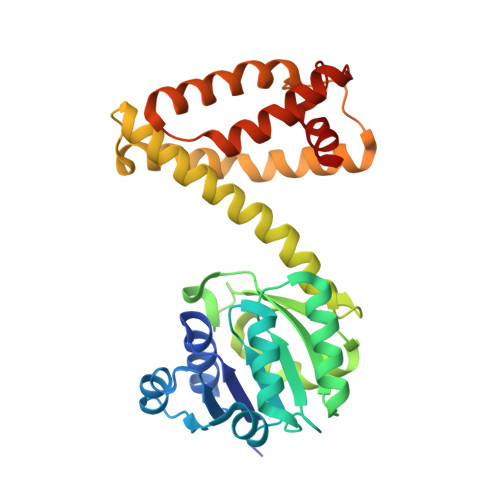
Structural Characterization of an S -enantioselective Imine Reductase from Mycobacterium Smegmatis .
Meyer, T., Zumbragel, N., Geerds, C., Groger, H., Niemann, H.H.(2020) Biomolecules 10
- PubMed: 32751900 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/biom10081130
- Primary Citation Related Structures:
6SMT - PubMed Abstract:
NADPH-dependent imine reductases (IREDs) are enzymes capable of enantioselectively reducing imines to chiral secondary amines, which represent important building blocks in the chemical and pharmaceutical industry. Since their discovery in 2011, many previously unknown IREDs have been identified, biochemically and structurally characterized and categorized into families. However, the catalytic mechanism and guiding principles for substrate specificity and stereoselectivity remain disputed. Herein, we describe the crystal structure of S -IRED- Ms from Mycobacterium smegmatis together with its cofactor NADPH. S -IRED- Ms belongs to the S -enantioselective superfamily 3 (SFam3) and is the first IRED from SFam3 to be structurally described. The data presented provide further evidence for the overall high degree of structural conservation between different IREDs of various superfamilies. We discuss the role of Asp170 in catalysis and the importance of hydrophobic amino acids in the active site for stereospecificity. Moreover, a separate entrance to the active site, potentially functioning according to a gatekeeping mechanism regulating access and, therefore, substrate specificity is described.
- Structural Biochemistry, Faculty of Chemistry, Bielefeld University, Universitätsstraße 25, 33615 Bielefeld, Germany.
Organizational Affiliation: