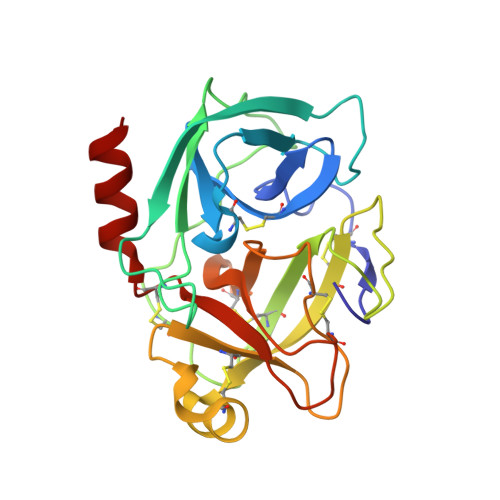
Structural Studies on the Inhibitory Binding Mode of Aromatic Coumarinic Esters to Human Kallikrein-Related Peptidase 7.
Hanke, S., Tindall, C.A., Pippel, J., Ulbricht, D., Pirotte, B., Reboud-Ravaux, M., Heiker, J.T., Strater, N.(2020) J Med Chem 63: 5723-5733
- PubMed: 32374603 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.9b01806
- Primary Citation Related Structures:
6SHH, 6SHI, 6SJU, 6Y4S - PubMed Abstract:
The serine protease kallikrein-related peptidase 7 (KLK7) is a member of the human tissue kallikreins. Its dysregulation leads to pathophysiological inflammatory processes in the skin. Furthermore, it plays a role in several types of cancer. For the treatment of KLK7-associated diseases, coumarinic esters have been developed as small-molecule enzyme inhibitors. To characterize the inhibition mode of these inhibitors, we analyzed structures of the inhibited protease by X-ray crystallography. Electron density shows the inhibitors covalently attached to His57 of the catalytic triad. This confirms the irreversible character of the inhibition process. Upon inhibitor binding, His57 undergoes an outward rotation; thus, the catalytic triad of the protease is disrupted. Besides, the halophenyl moiety of the inhibitor was absent in the final enzyme-inhibitor complex due to the hydrolysis of the ester linkage. With these results, we analyze the structural basis of KLK7 inhibition by the covalent attachment of aromatic coumarinic esters.
- Institute of Bioanalytical Chemistry, Center for Biotechnology and Biomedicine, Leipzig University, Deutscher Platz 5, 04103 Leipzig, Germany.
Organizational Affiliation: