Structure, Thermodynamics, and Kinetics of Plinabulin Binding to Two Tubulin Isotypes
La Sala, G., Olieric, N., Sharma, A.(2019) Chem 5
Experimental Data Snapshot
(2019) Chem 5
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Tubulin alpha-1B chain | 437 | Bos taurus | Mutation(s): 0 EC: 3.6.5 | 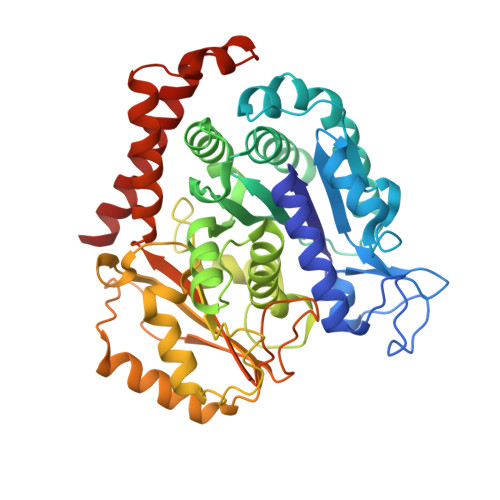 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P81947 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Tubulin beta-2B chain | 445 | Bos taurus | Mutation(s): 0 | 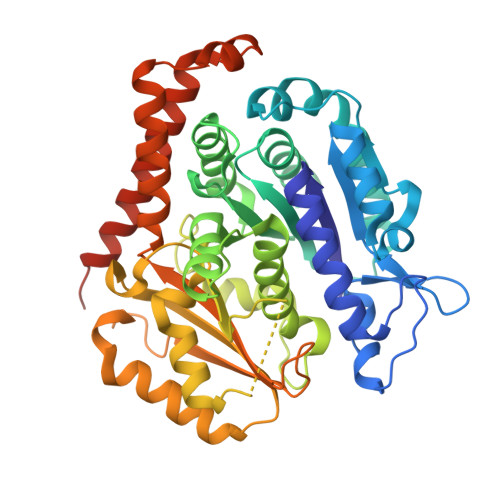 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q6B856 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Designed ankyrin repeat protein (DARPIN) D1 | C [auth F] | 155 | synthetic construct | Mutation(s): 0 | 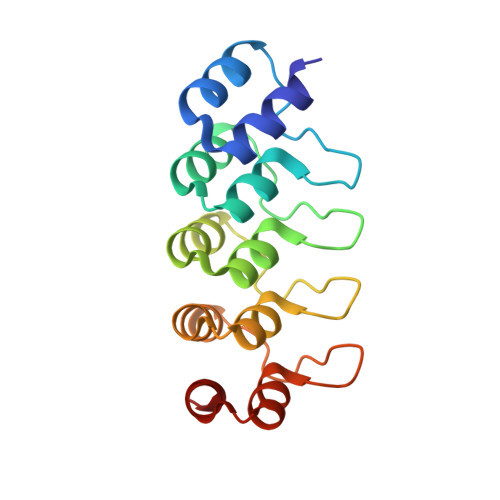 |
| Ligands 4 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| GTP Download:Ideal Coordinates CCD File | D [auth A] | GUANOSINE-5'-TRIPHOSPHATE C10 H16 N5 O14 P3 XKMLYUALXHKNFT-UUOKFMHZSA-N |  | ||
| GDP Download:Ideal Coordinates CCD File | G [auth B] | GUANOSINE-5'-DIPHOSPHATE C10 H15 N5 O11 P2 QGWNDRXFNXRZMB-UUOKFMHZSA-N |  | ||
| PN6 (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | F [auth B] | (3Z,6Z)-3-benzylidene-6-[(5-tert-butyl-1H-imidazol-4-yl)methylidene]piperazine-2,5-dione C19 H20 N4 O2 UNRCMCRRFYFGFX-TYPNBTCFSA-N |  | ||
| MG Download:Ideal Coordinates CCD File | E [auth A] | MAGNESIUM ION Mg JLVVSXFLKOJNIY-UHFFFAOYSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 73.565 | α = 90 |
| b = 91.351 | β = 96.851 |
| c = 83.221 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| XDS | data reduction |
| XSCALE | data scaling |
| PHASER | phasing |