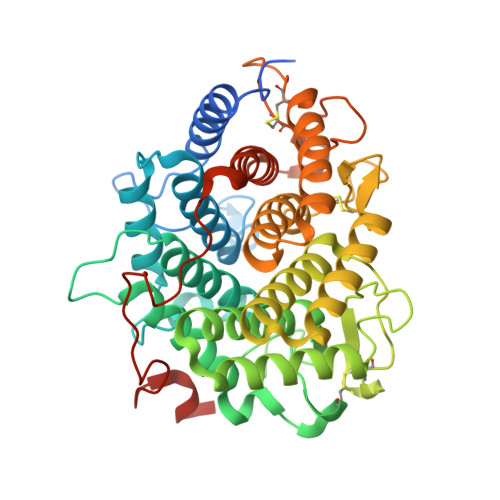
Structural base for the transfer of GPI-anchored glycoproteins into fungal cell walls.
Vogt, M.S., Schmitz, G.F., Varon Silva, D., Mosch, H.U., Essen, L.O.(2020) Proc Natl Acad Sci U S A 117: 22061-22067
- PubMed: 32839341 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2010661117
- Primary Citation Related Structures:
6RY0, 6RY1, 6RY2, 6RY5, 6RY6, 6RY7 - PubMed Abstract:
The correct distribution and trafficking of proteins are essential for all organisms. Eukaryotes evolved a sophisticated trafficking system which allows proteins to reach their destination within highly compartmentalized cells. One eukaryotic hallmark is the attachment of a glycosylphosphatidylinositol (GPI) anchor to C-terminal ω-peptides, which are used as a zip code to guide a subset of membrane-anchored proteins through the secretory pathway to the plasma membrane. In fungi, the final destination of many GPI-anchored proteins is their outermost compartment, the cell wall. Enzymes of the Dfg5 subfamily catalyze the essential transfer of GPI-anchored substrates from the plasma membrane to the cell wall and discriminate between plasma membrane-resident GPI-anchored proteins and those transferred to the cell wall (GPI-CWP). We solved the structure of Dfg5 from a filamentous fungus and used in crystallo glycan fragment screening to reassemble the GPI-core glycan in a U-shaped conformation within its binding pocket. The resulting model of the membrane-bound Dfg5•GPI-CWP complex is validated by molecular dynamics (MD) simulations and in vivo mutants in yeast. The latter show that impaired transfer of GPI-CWPs causes distorted cell-wall integrity as indicated by increased chitin levels. The structure of a Dfg5•β1,3-glycoside complex predicts transfer of GPI-CWP toward the nonreducing ends of acceptor glycans in the cell wall. In addition to our molecular model for Dfg5-mediated transglycosylation, we provide a rationale for how GPI-CWPs are specifically sorted toward the cell wall by using GPI-core glycan modifications.
- Department of Chemistry, Philipps-Universität, D-35032 Marburg, Germany.
Organizational Affiliation: