Evolved, Selective Erasers of Distinct Lysine Acylations.
Spinck, M., Neumann-Staubitz, P., Ecke, M., Gasper, R., Neumann, H.(2020) Angew Chem Int Ed Engl 59: 11142-11149
- PubMed: 32187803 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/anie.202002899
- Primary Citation Related Structures:
6RXJ, 6RXK, 6RXL, 6RXM, 6RXO, 6RXP, 6RXQ, 6RXR, 6RXS - PubMed Abstract:
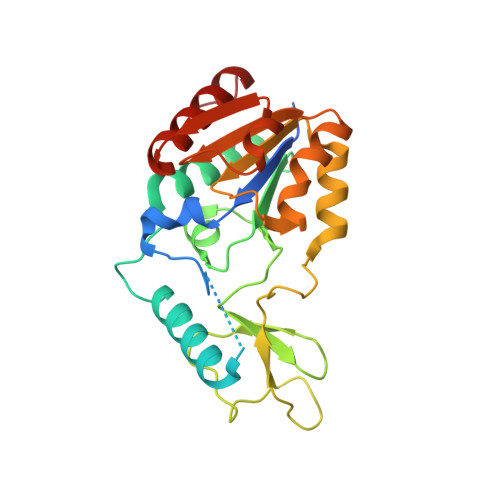
Lysine acylations, a family of diverse protein modifications varying in acyl-group length, charge, and saturation, are linked to many important physiological processes. Only a small set of substrate-promiscuous lysine acetyltransferases and deacetylases (KDACs) install and remove this vast variety of modifications. Engineered KDACs that remove only one type of acylation would help to dissect the different contributions of distinct acylations. We developed a bacterial selection system for the directed evolution of KDACs and identified variants up to 400 times more selective for butyryl-lysine compared to crotonyl-lysine. Structural analyses revealed that the enzyme adopts different conformational states depending on the type of acylation of the bound peptide. We used the butyryl-selective KDAC variant to shift the cellular acylation spectrum towards increased lysine crotonylation. These new enzymes will help in dissecting the roles of different lysine acylations in cell physiology.
- Department of Structural Biochemistry, Max-Planck-Institute of Molecular Physiology, Otto-Hahn-Strasse 11, 44227, Dortmund, Germany.
Organizational Affiliation: