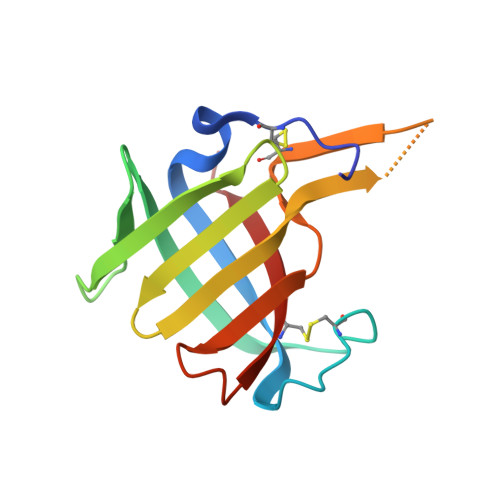
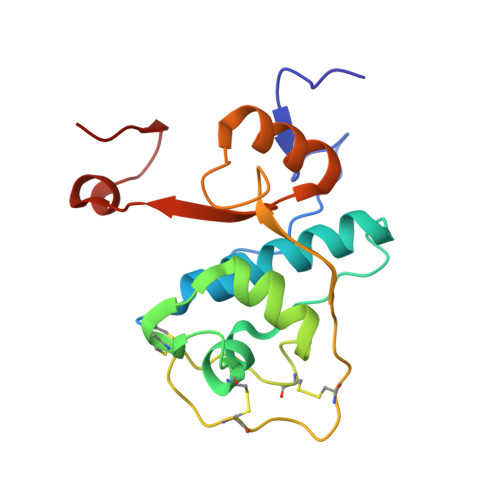
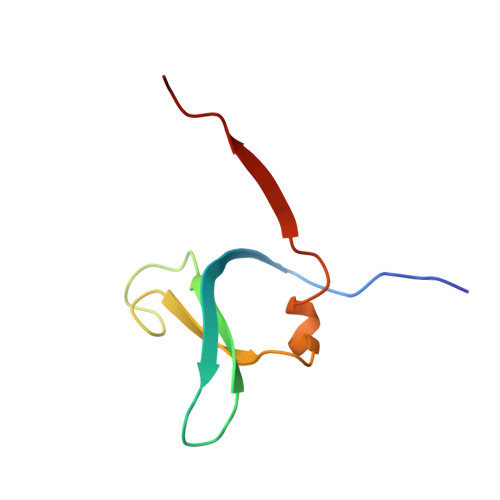
DPP1 Inhibitors: Exploring the Role of Water in the S2 Pocket of DPP1 with Substituted Pyrrolidines.
Kack, H., Doyle, K., Hughes, S.J., Bodnarchuk, M.S., Lonn, H., Van De Poel, A., Palmer, N.(2019) ACS Med Chem Lett 10: 1222-1227
- PubMed: 31413809 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acsmedchemlett.9b00261
- Primary Citation Related Structures:
6RN6, 6RN7, 6RN9, 6RNE, 6RNI - PubMed Abstract:
A series of pyrrolidine amino nitrile DPP1 inhibitors have been developed and characterized. The S2 pocket structure-activity relationship for these compounds shows significant gains in potency for DPP1 from interacting further with target residues and a network of water molecules in the binding pocket. Herein we describe the X-ray crystal structures of several of these compounds alongside an analysis of factors influencing the inhibitory potency toward DPP1 of which stabilization of the water network, demonstrated using Grand Canonical Monte Carlo simulations and free energy calculations, is attributed as a main factor.
- Structure, Biophysics and Fragments, Discovery Sciences, R&D, AstraZeneca, Gothenburg, Sweden.
Organizational Affiliation: