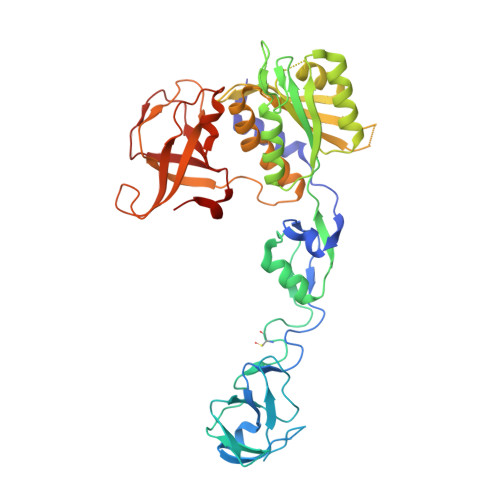
Insights into the Cnx1E catalyzed MPT-AMP hydrolysis.
Hercher, T.W., Krausze, J., Hoffmeister, S., Zwerschke, D., Lindel, T., Blankenfeldt, W., Mendel, R.R., Kruse, T.(2020) Biosci Rep 40
- PubMed: 31860061 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1042/BSR20191806
- Primary Citation Related Structures:
6RMS - PubMed Abstract:
Molybdenum insertases (Mo-insertases) catalyze the final step of molybdenum cofactor (Moco) biosynthesis, an evolutionary old and highly conserved multi-step pathway. In the first step of the pathway, GTP serves as substrate for the formation of cyclic pyranopterin monophosphate, which is subsequently converted into molybdopterin (MPT) in the second pathway step. In the following synthesis steps, MPT is adenylated yielding MPT-AMP that is subsequently used as substrate for enzyme catalyzed molybdate insertion. Molybdate insertion and MPT-AMP hydrolysis are catalyzed by the Mo-insertase E-domain. Earlier work reported a highly conserved aspartate residue to be essential for Mo-insertase functionality. In this work, we confirmed the mechanistic relevance of this residue for the Arabidopsis thaliana Mo-insertase Cnx1E. We found that the conservative substitution of Cnx1E residue Asp274 by Glu (D274E) leads to an arrest of MPT-AMP hydrolysis and hence to the accumulation of MPT-AMP. We further showed that the MPT-AMP accumulation goes in hand with the accumulation of molybdate. By crystallization and structure determination of the Cnx1E variant D274E, we identified the potential reason for the missing hydrolysis activity in the disorder of the region spanning amino acids 269 to 274. We reasoned that this is caused by the inability of a glutamate in position 274 to coordinate the octahedral Mg2+-water complex in the Cnx1E active site.
- TU Braunschweig, Institute of Plant Biology, Spielmannstrasse 7, 38106 Braunschweig, Germany.
Organizational Affiliation: