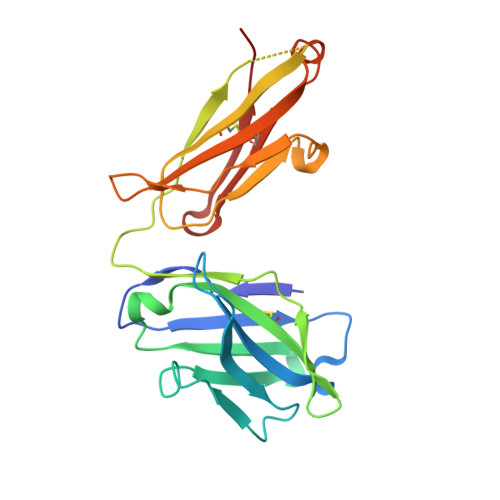
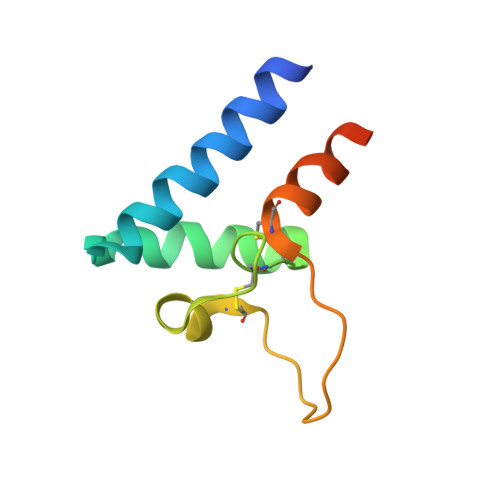
Structural basis of a homo-dimerization site in tetraspanin CD9 targeted by a melanoma patient-derived antibody
Neviani, V., Pos, W., Schotte, R., Wagner, K., Go, D.M., Fatmawati, C., Kedde, M., Claassen, Y.B., Kroon-Batenburg, L., Lutz, M., Verdegaal, E.M.E., van der Burg, S.H., Spits, H., Gros, P.To be published.