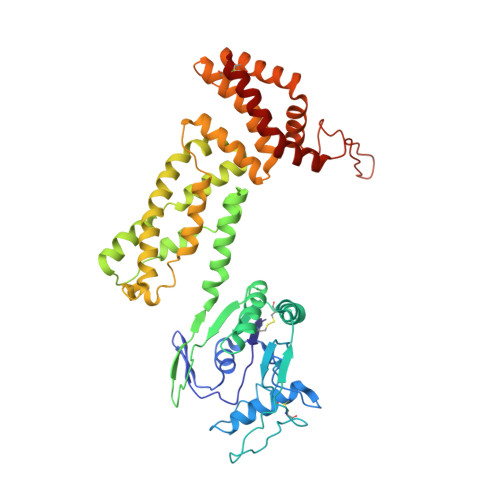
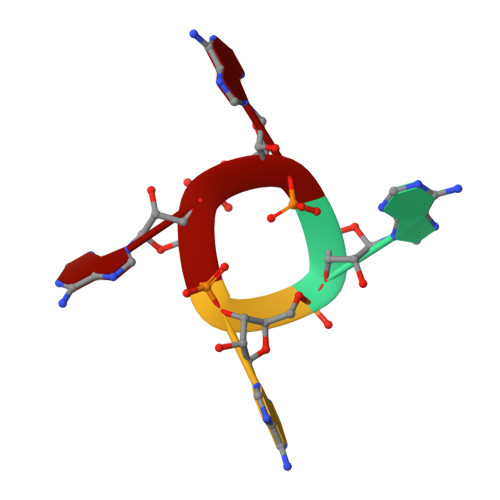
Structure of Csx1-cOA4complex reveals the basis of RNA decay in Type III-B CRISPR-Cas.
Molina, R., Stella, S., Feng, M., Sofos, N., Jauniskis, V., Pozdnyakova, I., Lopez-Mendez, B., She, Q., Montoya, G.(2019) Nat Commun 10: 4302-4302
- PubMed: 31541109 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-019-12244-z
- Primary Citation Related Structures:
6QZQ, 6QZT, 6R7B, 6R9R - PubMed Abstract:
Type III CRISPR-Cas multisubunit complexes cleave ssRNA and ssDNA. These activities promote the generation of cyclic oligoadenylate (cOA), which activates associated CRISPR-Cas RNases from the Csm/Csx families, triggering a massive RNA decay to provide immunity from genetic invaders. Here we present the structure of Sulfolobus islandicus (Sis) Csx1-cOA 4 complex revealing the allosteric activation of its RNase activity. SisCsx1 is a hexamer built by a trimer of dimers. Each dimer forms a cOA 4 binding site and a ssRNA catalytic pocket. cOA 4 undergoes a conformational change upon binding in the second messenger binding site activating ssRNA degradation in the catalytic pockets. Activation is transmitted in an allosteric manner through an intermediate HTH domain, which joins the cOA 4 and catalytic sites. The RNase functions in a sequential cooperative fashion, hydrolyzing phosphodiester bonds in 5'-C-C-3'. The degradation of cOA 4 by Ring nucleases deactivates SisCsx1, suggesting that this enzyme could be employed in biotechnological applications.
- Structural Molecular Biology Group, Novo Nordisk Foundation Centre for Protein Research, Faculty of Health and Medical Sciences University of Copenhagen, Blegdamsvej 3-B, Copenhagen, 2200, Denmark.
Organizational Affiliation: