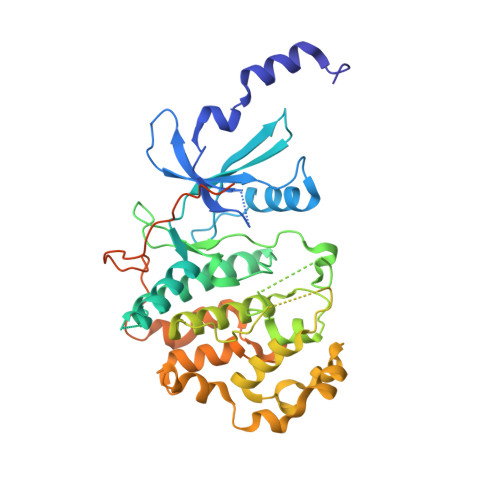
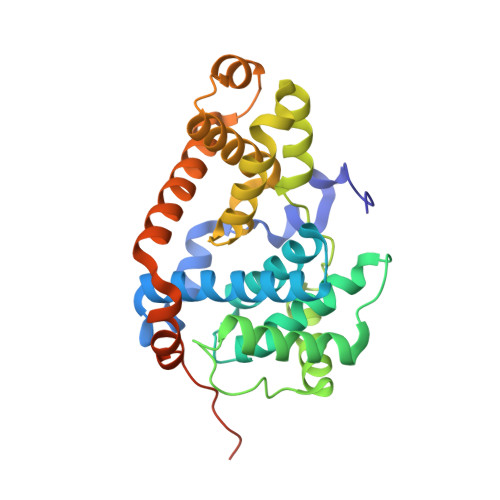
Selective and Potent CDK8/19 Inhibitors Enhance NK-Cell Activity and Promote Tumor Surveillance.
Hofmann, M.H., Mani, R., Engelhardt, H., Impagnatiello, M.A., Carotta, S., Kerenyi, M., Lorenzo-Herrero, S., Bottcher, J., Scharn, D., Arnhof, H., Zoephel, A., Schnitzer, R., Gerstberger, T., Sanderson, M.P., Rajgolikar, G., Goswami, S., Vasu, S., Ettmayer, P., Gonzalez, S., Pearson, M., McConnell, D.B., Kraut, N., Muthusamy, N., Moll, J.(2020) Mol Cancer Ther 19: 1018-1030
- PubMed: 32024684 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1158/1535-7163.MCT-19-0789
- Primary Citation Related Structures:
6QTG, 6QTJ, 6R3S - PubMed Abstract:
Natural killer (NK) cells play a pivotal role in controlling cancer. Multiple extracellular receptors and internal signaling nodes tightly regulate NK activation. Cyclin-dependent kinases of the mediator complex (CDK8 and CDK19) were described as a signaling intermediates in NK cells. Here, we report for the first time the development and use of CDK8/19 inhibitors to suppress phosphorylation of STAT1 S727 in NK cells and to augment the production of the cytolytic molecules perforin and granzyme B (GZMB). Functionally, this resulted in enhanced NK-cell-mediated lysis of primary leukemia cells. Treatment with the CDK8/19 inhibitor BI-1347 increased the response rate and survival of mice bearing melanoma and breast cancer xenografts. In addition, CDK8/19 inhibition augmented the antitumoral activity of anti-PD-1 antibody and SMAC mimetic therapy, both agents that promote T-cell-mediated antitumor immunity. Treatment with the SMAC mimetic compound BI-8382 resulted in an increased number of NK cells infiltrating EMT6 tumors. Combination of the CDK8/19 inhibitor BI-1347, which augments the amount of degranulation enzymes, with the SMAC mimetic BI-8382 resulted in increased survival of mice carrying the EMT6 breast cancer model. The observed survival benefit was dependent on an intermittent treatment schedule of BI-1347, suggesting the importance of circumventing a hyporesponsive state of NK cells. These results suggest that CDK8/19 inhibitors can be combined with modulators of the adaptive immune system to inhibit the growth of solid tumors, independent of their activity on cancer cells, but rather through promoting NK-cell function.
- Boehringer Ingelheim RCV GmbH & Co KG, Vienna, Austria. marco.hofmann@boehringer-ingelheim.com.
Organizational Affiliation: