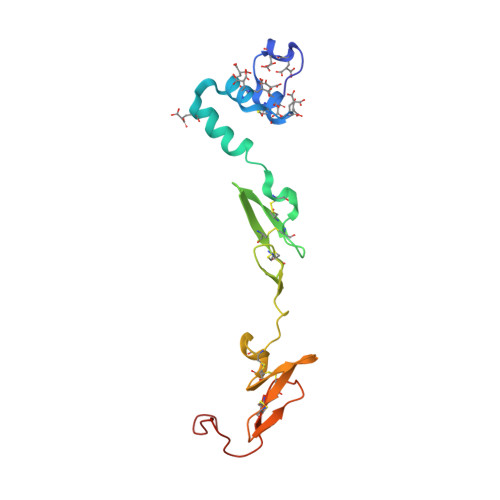
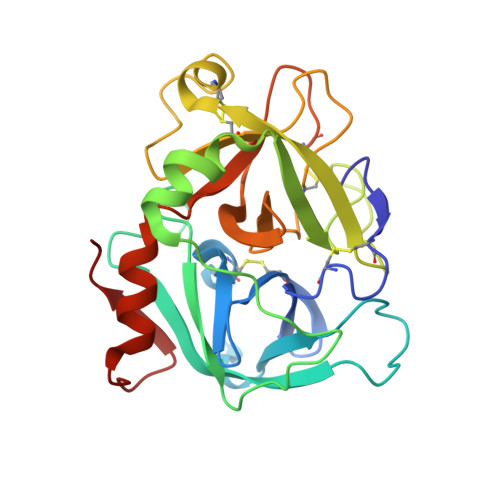
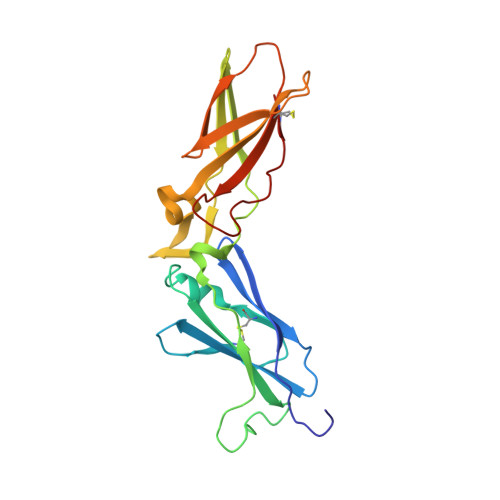
Beating tissue factor at its own game: Design and properties of a soluble tissue factor-independent coagulation factor VIIa.
Sorensen, A.B., Tuneew, I., Svensson, L.A., Persson, E., Ostergaard, H., Overgaard, M.T., Olsen, O.H., Gandhi, P.S.(2020) J Biological Chem 295: 517-528
- PubMed: 31801825 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA119.009183
- Primary Citation Related Structures:
6R2W - PubMed Abstract:
Two decades of research have uncovered the mechanism by which the complex of tissue factor (TF) and the plasma serine protease factor VIIa (FVIIa) mediates the initiation of blood coagulation. Membrane-anchored TF directly interacts with substrates and induces allosteric effects in the protease domain of FVIIa. These properties are also recapitulated by the soluble ectodomain of TF (sTF). At least two interdependent allosteric activation pathways originate at the FVIIa:sTF interface are proposed to enhance FVIIa activity upon sTF binding. Here, we sought to engineer an sTF-independent FVIIa variant by stabilizing both proposed pathways, with one pathway terminating at segment 215-217 in the activation domain and the other pathway terminating at the N terminus insertion site. To stabilize segment 215-217, we replaced the flexible 170 loop of FVIIa with the more rigid 170 loop from trypsin and combined it with an L163V substitution (FVIIa-VY T ). The FVIIa-VYT variant exhibited 60-fold higher amidolytic activity than FVIIa, and displayed similar FX activation and antithrombin inhibition kinetics to the FVIIa.sTF complex. The sTF-independent activity of FVIIa-VY T was partly mediated by an increase in the N terminus insertion and, as shown by X-ray crystallography, partly by Tyr-172 inserting into a cavity in the activation domain stabilizing the S1 substrate-binding pocket. The combination with L163V likely drove additional changes in a delicate hydrogen-bonding network that further stabilized S1-S3 sites. In summary, we report the first FVIIa variant that is catalytically independent of sTF and provide evidence supporting the existence of two TF-mediated allosteric activation pathways.
- Global Research, Novo Nordisk A/S, DK-2760 Måløv, Denmark; Department of Chemistry and Bioscience, Aalborg University, DK-9220 Aalborg, Denmark. Electronic address: abs@evaxion-biotech.com.
Organizational Affiliation: