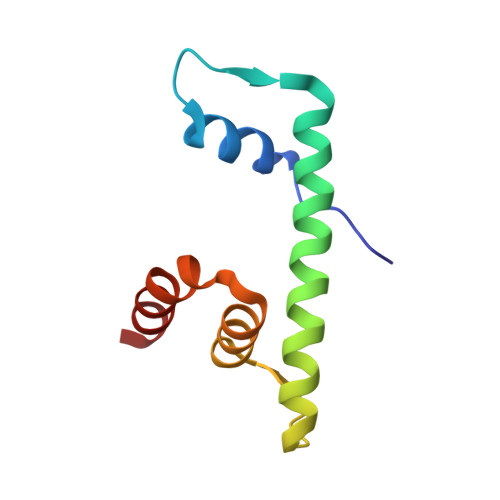
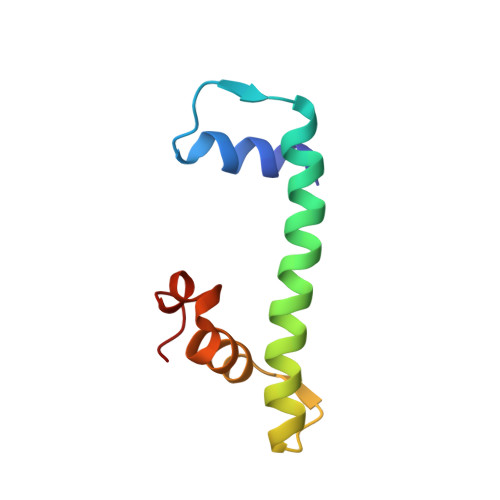
Structural determinants for NF-Y subunit organization and NF-Y/DNA association in plants.
Chaves-Sanjuan, A., Gnesutta, N., Gobbini, A., Martignago, D., Bernardini, A., Fornara, F., Mantovani, R., Nardini, M.(2021) Plant J 105: 49-61
- PubMed: 33098724 Search on PubMed
- DOI: https://doi.org/10.1111/tpj.15038
- Primary Citation Related Structures:
6R0L, 6R0M, 6R0N, 6R2V - PubMed Abstract:
NF-Y transcription factor comprises three subunits: NF-YA, NF-YB and NF-YC. NF-YB and NF-YC dimerize through their histone fold domain (HFD), which can bind DNA in a non-sequence-specific fashion while serving as a scaffold for NF-YA trimerization. Upon trimerization, NF-YA specifically recognizes the CCAAT box sequence on promoters and enhancers. In plants, each NF-Y subunit is encoded by several genes giving rise to hundreds of potential heterotrimeric combinations. In addition, plant NF-YBs and NF-YCs interact with other protein partners to recognize a plethora of genomic motifs, as the CCT protein family that binds CORE sites. The NF-Y subunit organization and its DNA-binding properties, together with the NF-Y HFD capacity to adapt different protein modules, represent plant-specific features that play a key role in development, growth and reproduction. Despite their relevance, these features are still poorly understood at the molecular level. Here, we present the structures of Arabidopsis and rice NF-YB/NF-YC dimers, and of an Arabidopsis NF-Y trimer in complex with the FT CCAAT box, together with biochemical data on NF-Y mutants. The dimeric structures identify the key residues for NF-Y HFD stabilization. The NF-Y/DNA structure and the mutation experiments shed light on HFD trimerization interface properties and the NF-YA sequence appetite for the bases flanking the CCAAT motif. These data explain the logic of plant NF-Y gene expansion: the trimerization adaptability and the flexible DNA-binding rules serve the scopes of accommodating the large number of NF-YAs, CCTs and possibly other NF-Y HFD binding partners and a diverse audience of genomic motifs.
- Dipartimento di Bioscienze, Università degli Studi di Milano, Via Celoria 26, Milano, 20133, Italy.
Organizational Affiliation: