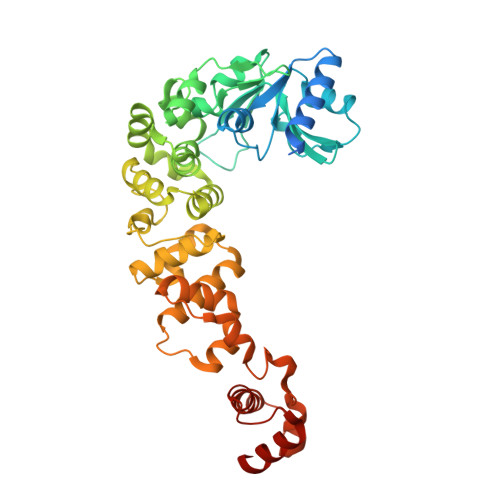
CCA-addition in the cold: Structural characterization of the psychrophilic CCA-adding enzyme from the permafrost bacterium Planococcus halocryophilus
de Wijn, R., Rollet, K., Ernst, F.G., Wellner, K., Betat, H., Morl, M., Sauter, C.(2021) Comput Struct Biotechnol J