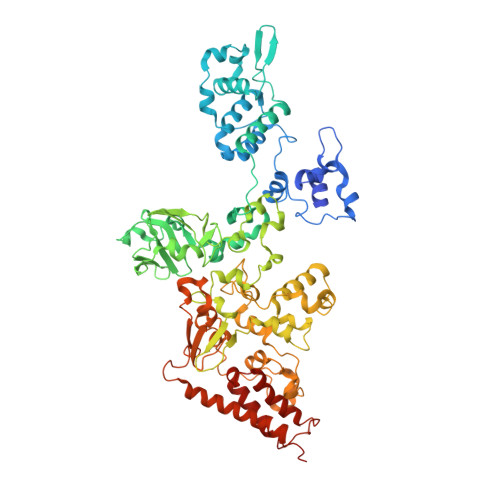
Extracellular Fe(III) reductase structure reveals a modular organization enabling S-layer insertion and electron transfer to insoluble substrates
Tikhonova, T.V., Osipov, E.M., Dergousova, N.I., Boyko, K.M., Elizarov, I.M., Gavrilov, S.N., Khrenova, M.G., Robb, F.T., Solovieva, A.Y., Bonch-Osmolovskaya, E.A., Popov, V.O.(2023) Structure