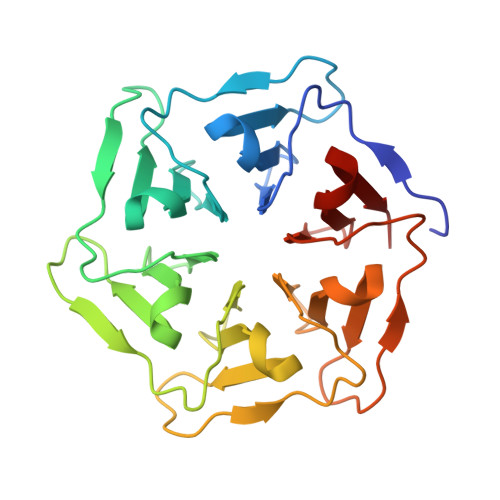
Hybrid assemblies of a symmetric designer protein and polyoxometalates with matching symmetry.
Vandebroek, L., Noguchi, H., Kamata, K., Tame, J.R.H., Van Meervelt, L., Parac-Vogt, T.N., Voet, A.R.D.(2020) Chem Commun (Camb) 56: 11601-11604
- PubMed: 32869783 Search on PubMed
- DOI: https://doi.org/10.1039/d0cc05071g
- Primary Citation Related Structures:
6QSD, 6QSE, 6QSF, 6QSG, 6QSH - PubMed Abstract:
Novel bioinorganic hybrid materials based on proteins and inorganic clusters have enormous potential for the development of hybrid catalysts that synergistically combine properties of both materials. Here we report the creation of hybrid assemblies between a computationally designed symmetrical protein Pizza6-S and different polyoxometalates with matching symmetry: the tellurotungstic Anderson-Evans (Na6[TeW6O24]·22H2O) (TEW); Keggin (H4[SiW12O40]·6H2O) (STA); and 1 : 2 CeIII-substituted Keggin (K11[CeIII[PW11O39]2]·20H2O) (Ce-K). This results in the formation of complexes with clearly defined stoichiometries in solution. Crystal structures validate the complexes as building blocks for the formation of larger assemblies.
- Laboratory for Bioinorganic Chemistry, KU Leuven Department of Chemistry, Celestijnenlaan 200F, 3001 Leuven, Belgium and Biomolecular Architecture, KU Leuven Department of Chemistry, Celestijnenlaan 200F, 3001 Leuven, Belgium.
Organizational Affiliation: