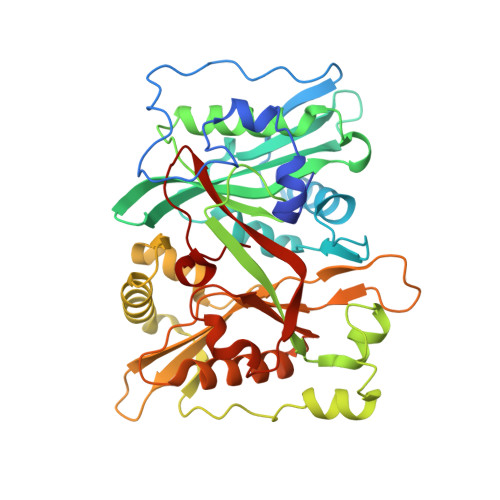
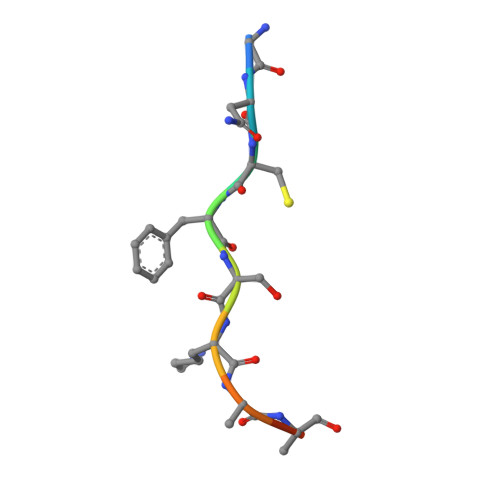
High-resolution snapshots of human N-myristoyltransferase in action illuminate a mechanism promoting N-terminal Lys and Gly myristoylation.
Dian, C., Perez-Dorado, I., Riviere, F., Asensio, T., Legrand, P., Ritzefeld, M., Shen, M., Cota, E., Meinnel, T., Tate, E.W., Giglione, C.(2020) Nat Commun 11: 1132-1132
- PubMed: 32111831 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-020-14847-3
- Primary Citation Related Structures:
6EHJ, 6QRM, 6SJZ, 6SK2, 6SK3, 6SK8, 6SKJ - PubMed Abstract:
The promising drug target N-myristoyltransferase (NMT) catalyses an essential protein modification thought to occur exclusively at N-terminal glycines (Gly). Here, we present high-resolution human NMT1 structures co-crystallised with reactive cognate lipid and peptide substrates, revealing high-resolution snapshots of the entire catalytic mechanism from the initial to final reaction states. Structural comparisons, together with biochemical analysis, provide unforeseen details about how NMT1 reaches a catalytically competent conformation in which the reactive groups are brought into close proximity to enable catalysis. We demonstrate that this mechanism further supports efficient and unprecedented myristoylation of an N-terminal lysine side chain, providing evidence that NMT acts both as N-terminal-lysine and glycine myristoyltransferase.
- Université Paris-Saclay, CEA, CNRS, Institute for Integrative Biology of the Cell (I2BC), Gif-sur-Yvette, 91198, France.
Organizational Affiliation: