Serine bacteriolytic protease L1 of Lysobacter sp. XL1 complexed with protease inhibitor AEBSF: features of interaction
Kudryakova, I., Gabdulkhakov, A., Tishchenko, S., Afoshin, A., Vasilyeva, N.(2019) Process Biochem
Experimental Data Snapshot
Starting Model: experimental
View more details
(2019) Process Biochem
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Lytic endopeptidase preproenzyme | 199 | Lysobacter sp. XL1 | Mutation(s): 0 Gene Names: alpA | 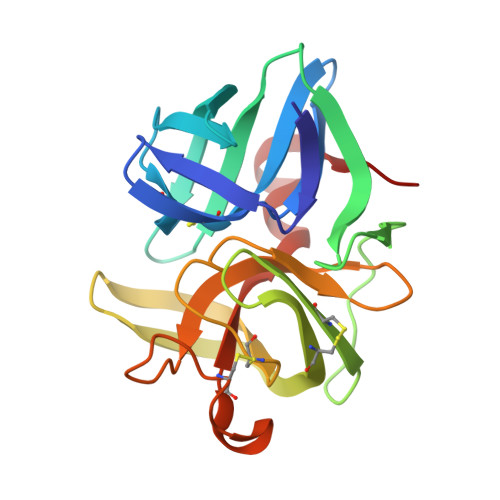 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | D2K8B3 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 7 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| AES Download:Ideal Coordinates CCD File | BA [auth D], H [auth A], M [auth B], U [auth C] | 4-(2-AMINOETHYL)BENZENESULFONYL FLUORIDE C8 H10 F N O2 S MGSKVZWGBWPBTF-UHFFFAOYSA-N |  | ||
| JAT Download:Ideal Coordinates CCD File | CA [auth D], N [auth B], V [auth C] | 4-(2-azanylethyl)benzenesulfonic acid C8 H11 N O3 S RYBFWJVHZIKGQJ-UHFFFAOYSA-N |  | ||
| PGE Download:Ideal Coordinates CCD File | K [auth A] | TRIETHYLENE GLYCOL C6 H14 O4 ZIBGPFATKBEMQZ-UHFFFAOYSA-N |  | ||
| PEG Download:Ideal Coordinates CCD File | Q [auth B], R [auth B], Y [auth C] | DI(HYDROXYETHYL)ETHER C4 H10 O3 MTHSVFCYNBDYFN-UHFFFAOYSA-N |  | ||
| SO4 Download:Ideal Coordinates CCD File | AA [auth D] E [auth A] F [auth A] G [auth A] L [auth B] | SULFATE ION O4 S QAOWNCQODCNURD-UHFFFAOYSA-L |  | ||
| GOL Download:Ideal Coordinates CCD File | DA [auth D], I [auth A], J [auth A], P [auth B], X [auth C] | GLYCEROL C3 H8 O3 PEDCQBHIVMGVHV-UHFFFAOYSA-N |  | ||
| CL Download:Ideal Coordinates CCD File | O [auth B], W [auth C] | CHLORIDE ION Cl VEXZGXHMUGYJMC-UHFFFAOYSA-M |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 41.855 | α = 90 |
| b = 122.553 | β = 98.64 |
| c = 78.988 | γ = 90 |
| Software Name | Purpose |
|---|---|
| PHENIX | refinement |
| XDS | data reduction |
| XSCALE | data scaling |
| PHASER | phasing |