The role of the light chain in the structure and binding activity of two cattle antibodies that neutralize bovine respiratory syncytial virus.
Ren, J., Nettleship, J.E., Harris, G., Mwangi, W., Rhaman, N., Grant, C., Kotecha, A., Fry, E., Charleston, B., Stuart, D.I., Hammond, J., Owens, R.J.(2019) Mol Immunol 112: 123-130
- PubMed: 31100550 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molimm.2019.04.026
- Primary Citation Related Structures:
6QN7, 6QN8, 6QN9, 6QNA - PubMed Abstract:
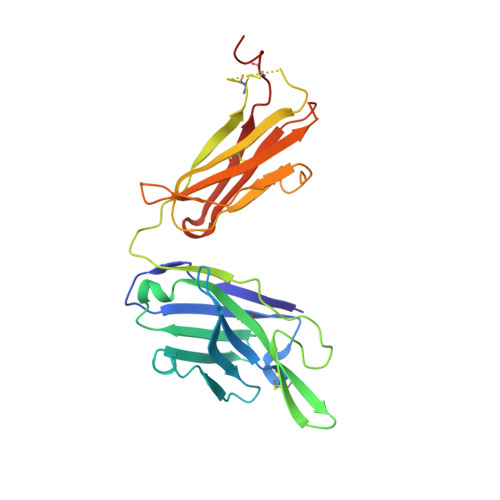
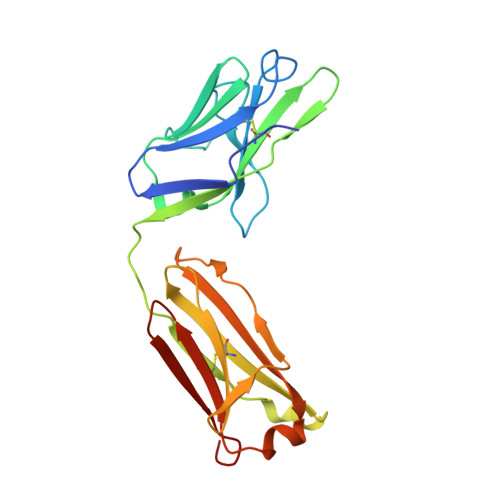
Cattle antibodies have unusually long CDR3 loops in their heavy chains (HCs), and limited light chain (LC) diversity, raising the question of whether these mask the effect of LC variation on antigen recognition. We have investigated the role of the LC in the structure and activity of two neutralizing cattle antibodies (B4 and B13) that bind the F protein of bovine respiratory syncytial virus (bRSV). Recombinant Fab fragments of B4 and B13 bound bRSV infected cells and showed similar affinities for purified bRSV F protein. Exchanging the LCs between the Fab fragments produced hybrid Fabs: B13* (B13 HC/B4 LC) and B4* (B4 HC/B13 LC). The affinity of B13* to the F protein was found to be two-fold lower than B13 whilst the binding affinity of B4* was reduced at least a hundred-fold compared to B4 such that it no longer bound to bRSV infected cells. Comparison of the structures of B4 and B13 with their LC exchanged counterparts B4* and B13* showed that paratope of the HC variable domain (VH) of B4 was disrupted on pairing with the B13 LC, consistent with the loss of binding activity. By contrast, B13 H3 adopts a similar conformation when paired with either B13 or B4 LCs. These observations confirm the expected key role of the extended H3 loop in antigen-binding by cattle antibodies but also show that the quaternary LC/HC subunit interaction can be crucial for its presentation and thus the LC variable domain (VL) is also important for antigen recognition.
- The Division of Structural Biology, Henry Wellcome Building for Genomic Medicine, Roosevelt Drive, Oxford, OX3 7BN, UK.
Organizational Affiliation: