Novel Thienopyrimidine Inhibitors of Leishmania N -Myristoyltransferase with On-Target Activity in Intracellular Amastigotes.
Bell, A.S., Yu, Z., Hutton, J.A., Wright, M.H., Brannigan, J.A., Paape, D., Roberts, S.M., Sutherell, C.L., Ritzefeld, M., Wilkinson, A.J., Smith, D.F., Leatherbarrow, R.J., Tate, E.W.(2020) J Med Chem 63: 7740-7765
- PubMed: 32575985 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.jmedchem.0c00570
- Primary Citation Related Structures:
6QD9, 6QDA, 6QDB, 6QDC, 6QDD, 6QDE, 6QDF, 6QDG, 6QDH - PubMed Abstract:
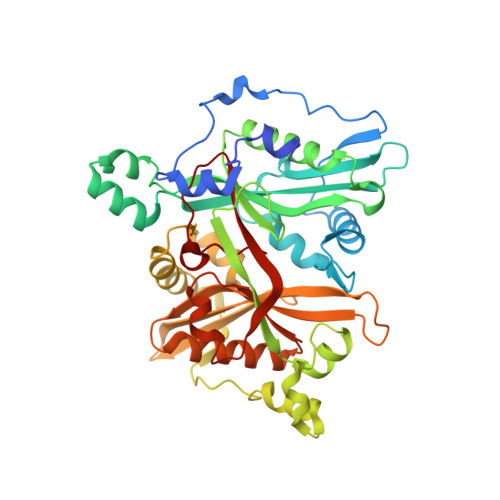
The leishmaniases, caused by Leishmania species of protozoan parasites, are neglected tropical diseases with millions of cases worldwide. Current therapeutic approaches are limited by toxicity, resistance, and cost. N -Myristoyltransferase (NMT), an enzyme ubiquitous and essential in all eukaryotes, has been validated via genetic and pharmacological methods as a promising anti-leishmanial target. Here we describe a comprehensive structure-activity relationship (SAR) study of a thienopyrimidine series previously identified in a high-throughput screen against Leishmania NMT, across 68 compounds in enzyme- and cell-based assay formats. Using a chemical tagging target engagement biomarker assay, we identify the first inhibitor in this series with on-target NMT activity in leishmania parasites. Furthermore, crystal structure analyses of 12 derivatives in complex with Leishmania major NMT revealed key factors important for future structure-guided optimization delivering IMP-105 ( 43 ), a compound with modest activity against Leishmania donovani intracellular amastigotes and excellent selectivity (>660-fold) for Leishmania NMT over human NMTs.
- Department of Chemistry, Imperial College London, Molecular Sciences Research Hub, London, U.K. W12 0BZ.
Organizational Affiliation: