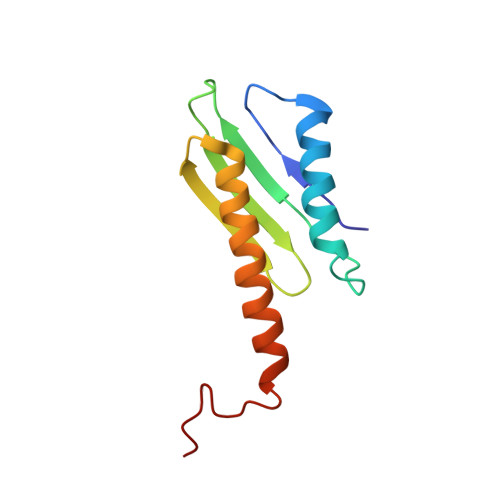
Solution structure of the N-terminal domain of the Staphylococcus aureus hibernation promoting factor.
Usachev, K.S., Validov, S.Z., Khusainov, I.S., Varfolomeev, A.A., Klochkov, V.V., Aganov, A.V., Yusupov, M.M.(2019) J Biomol NMR 73: 223-227
- PubMed: 31165320 Search on PubMed
- DOI: https://doi.org/10.1007/s10858-019-00254-4
- Primary Citation Related Structures:
6QBZ - PubMed Abstract:
Staphylococcus aureus hibernation promoting factor (SaHPF) is a 22,2 kDa protein which plays a crucial role in 100S Staphylococcus aureus ribosome formation during stress. SaHPF consists of N-terminal domain (NTD) that prevents proteins synthesis by binding to the 30S subunit at the P- and A-sites, connected through a flexible linker with a C-terminal domain (CTD) that keeps ribosomes in 100S form via homodimerization. Recently obtained 100S ribosome structure of S. aureus by cryo-EM shown that SaHPF-NTD bound to the ribosome active sites, however due to the absence of SaHPF-NTD structure it was modeled by homology with the E. coli hibernation factors HPF and YfiA. In present paper we have determined the solution structure of SaHPF-NTD by high-resolution NMR spectroscopy which allows us to increase structural knowledge about HPF structure from S. aureus.
- Laboratory of Structural Biology, Institute of Fundamental Medicine and Biology, Kazan Federal University, 18 Kremlevskaya, Kazan, 420008, Russian Federation.
Organizational Affiliation: