Structure Elucidation of Helical Aromatic Foldamer-Protein Complexes with Large Contact Surface Areas.
Reddy, P.S., Langlois d'Estaintot, B., Granier, T., Mackereth, C.D., Fischer, L., Huc, I.(2019) Chemistry 25: 11042-11047
- PubMed: 31257622 Search on PubMed
- DOI: https://doi.org/10.1002/chem.201902942
- Primary Citation Related Structures:
6HZX, 6Q9T - PubMed Abstract:
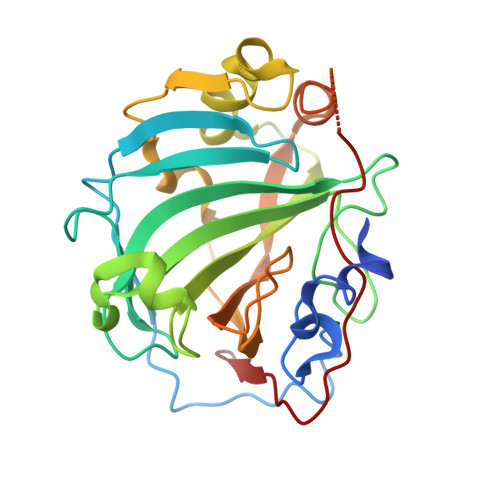
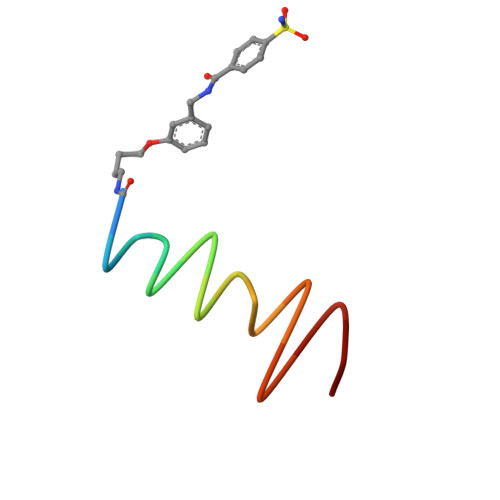
The development of large synthetic ligands could be useful to target the sizeable surface areas involved in protein-protein interactions. Herein, we present long helical aromatic oligoamide foldamers bearing proteinogenic side chains that cover up to 450 Å 2 of the human carbonic anhydrase II (HCA) surface. The foldamers are composed of aminoquinolinecarboxylic acids bearing proteinogenic side chains and of more flexible aminomethyl-pyridinecarboxylic acids that enhance helix handedness dynamics. Crystal structures of HCA-foldamer complexes were obtained with a 9- and a 14-mer both showing extensive protein-foldamer hydrophobic contacts. In addition, foldamer-foldamer interactions seem to be prevalent in the crystal packing, leading to the peculiar formation of an HCA superhelix wound around a rod of stacked foldamers. Solution studies confirm the positioning of the foldamer at the protein surface as well as a dimerization of the complexes.
- CBMN (UMR5248), Univ. Bordeaux-CNRS-INP, Institut Européen de Chimie et Biologie, 2 rue Escarpit, 33600, Pessac, France.
Organizational Affiliation: