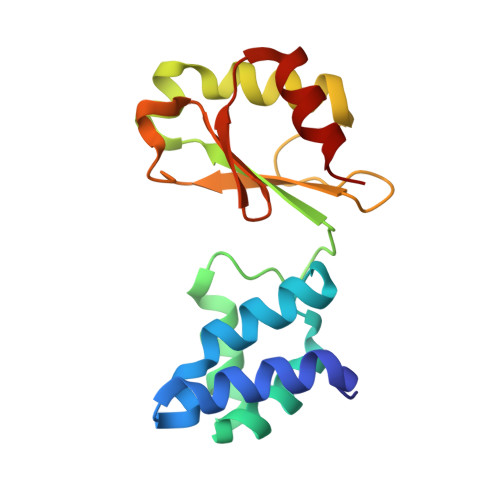
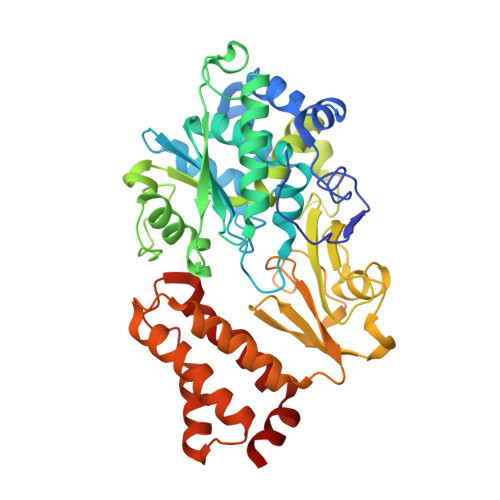
A mechanism to prevent production of reactive oxygen species by Escherichia coli respiratory complex I.
Schulte, M., Frick, K., Gnandt, E., Jurkovic, S., Burschel, S., Labatzke, R., Aierstock, K., Fiegen, D., Wohlwend, D., Gerhardt, S., Einsle, O., Friedrich, T.(2019) Nat Commun 10: 2551-2551
- PubMed: 31186428 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-019-10429-0
- Primary Citation Related Structures:
6HL2, 6HL3, 6HL4, 6HLA, 6HLI, 6HLJ, 6HLM, 6Q9C, 6Q9G, 6Q9J, 6Q9K, 6R7P - PubMed Abstract:
Respiratory complex I plays a central role in cellular energy metabolism coupling NADH oxidation to proton translocation. In humans its dysfunction is associated with degenerative diseases. Here we report the structure of the electron input part of Aquifex aeolicus complex I at up to 1.8 Å resolution with bound substrates in the reduced and oxidized states. The redox states differ by the flip of a peptide bond close to the NADH binding site. The orientation of this peptide bond is determined by the reduction state of the nearby [Fe-S] cluster N1a. Fixation of the peptide bond by site-directed mutagenesis led to an inactivation of electron transfer and a decreased reactive oxygen species (ROS) production. We suggest the redox-gated peptide flip to represent a previously unrecognized molecular switch synchronizing NADH oxidation in response to the redox state of the complex as part of an intramolecular feed-back mechanism to prevent ROS production.
- Institut für Biochemie, Albert-Ludwigs-Universität Freiburg, Albertstr. 21, 79104, Freiburg, Germany.
Organizational Affiliation: