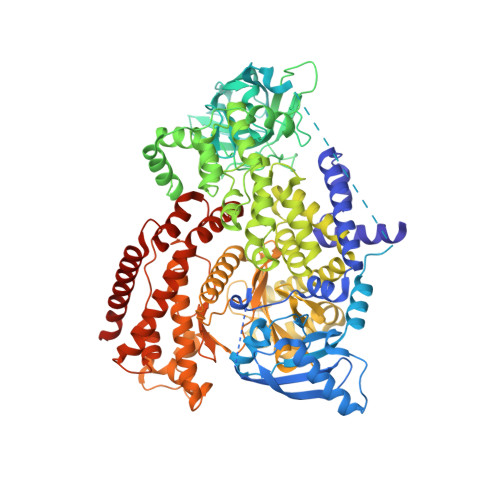
Discovery of Potent, Efficient, and Selective Inhibitors of Phosphoinositide 3-Kinase delta through a Deconstruction and Regrowth Approach.
Barton, N., Convery, M., Cooper, A.W.J., Down, K., Hamblin, J.N., Inglis, G., Peace, S., Rowedder, J., Rowland, P., Taylor, J.A., Wellaway, N.(2018) J Med Chem 61: 11061-11073
- PubMed: 30532965 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.8b01556
- Primary Citation Related Structures:
6Q6Y, 6Q73, 6Q74 - PubMed Abstract:
A deconstruction of previously reported phosphoinositide 3-kinase δ (PI3Kδ) inhibitors and subsequent regrowth led to the identification of a privileged fragment for PI3Kδ, which was exploited to deliver a potent, efficient, and selective lead series with a novel binding mode observed in the PI3Kδ crystal structure.
- GlaxoSmithKline R&D, Medicines Research Centre , Gunnels Wood Road , SG1 2NY Stevenage , U.K.
Organizational Affiliation: