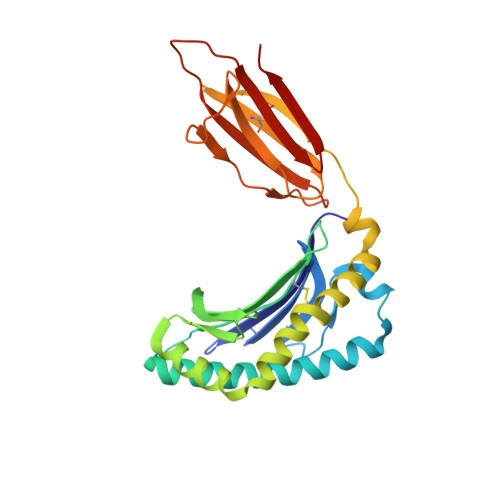
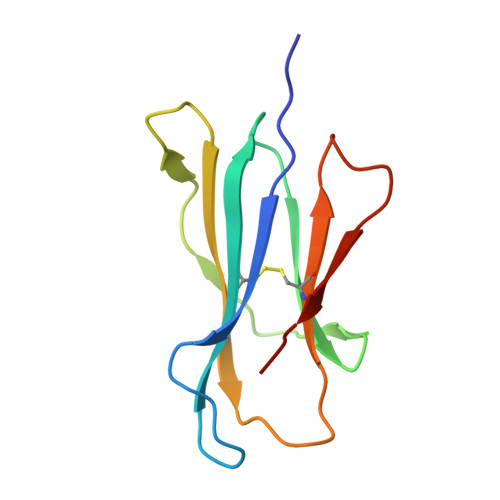
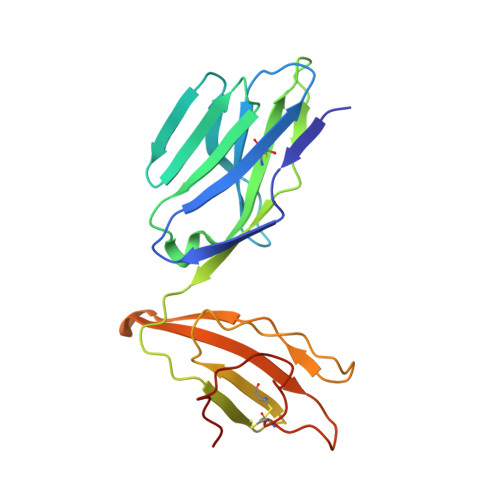
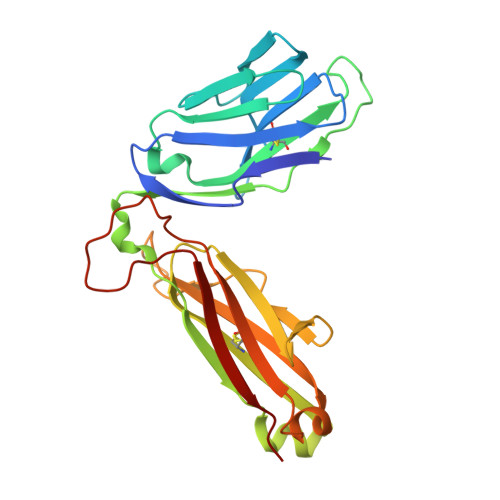
Ligand-dependent downregulation of MR1 cell surface expression.
Salio, M., Awad, W., Veerapen, N., Gonzalez-Lopez, C., Kulicke, C., Waithe, D., Martens, A.W.J., Lewinsohn, D.M., Hobrath, J.V., Cox, L.R., Rossjohn, J., Besra, G.S., Cerundolo, V.(2020) Proc Natl Acad Sci U S A 117: 10465-10475
- PubMed: 32341160 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2003136117
- Primary Citation Related Structures:
6PVC, 6PVD - PubMed Abstract:
The antigen-presenting molecule MR1 presents riboflavin-based metabolites to Mucosal-Associated Invariant T (MAIT) cells. While MR1 egress to the cell surface is ligand-dependent, the ability of small-molecule ligands to impact on MR1 cellular trafficking remains unknown. Arising from an in silico screen of the MR1 ligand-binding pocket, we identify one ligand, 3-([2,6-dioxo-1,2,3,6-tetrahydropyrimidin-4-yl]formamido)propanoic acid, DB28, as well as an analog, methyl 3-([2,6-dioxo-1,2,3,6-tetrahydropyrimidin-4-yl]formamido)propanoate, NV18.1, that down-regulate MR1 from the cell surface and retain MR1 molecules in the endoplasmic reticulum (ER) in an immature form. DB28 and NV18.1 compete with the known MR1 ligands, 5-OP-RU and acetyl-6-FP, for MR1 binding and inhibit MR1-dependent MAIT cell activation. Crystal structures of the MAIT T cell receptor (TCR) complexed with MR1-DB28 and MR1-NV18.1, show that these two ligands reside within the A'-pocket of MR1. Neither ligand forms a Schiff base with MR1 molecules; both are nevertheless sequestered by a network of hydrophobic and polar contacts. Accordingly, we define a class of compounds that inhibits MR1 cellular trafficking.
- MRC Human Immunology Unit, MRC Weatherall Institute of Molecular Medicine, University of Oxford, Oxford OX3 9DS, United Kingdom; mariolina.salio@imm.ox.ac.uk g.besra@bham.ac.uk.
Organizational Affiliation: