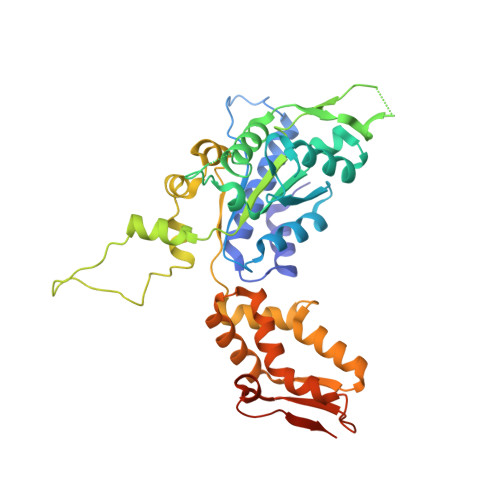
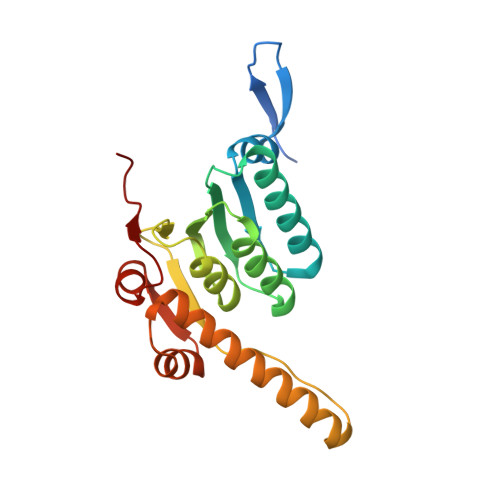
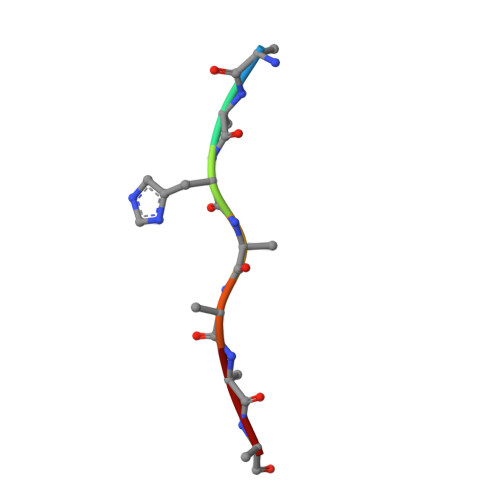
Structures of the ATP-fueled ClpXP proteolytic machine bound to protein substrate.
Fei, X., Bell, T.A., Jenni, S., Stinson, B.M., Baker, T.A., Harrison, S.C., Sauer, R.T.(2020) Elife 9
- PubMed: 32108573 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.52774
- Primary Citation Related Structures:
6PO1, 6PO3, 6POD, 6POS, 6PP5, 6PP6, 6PP7, 6PP8, 6PPE - PubMed Abstract:
ClpXP is an ATP-dependent protease in which the ClpX AAA+ motor binds, unfolds, and translocates specific protein substrates into the degradation chamber of ClpP. We present cryo-EM studies of the E. coli enzyme that show how asymmetric hexameric rings of ClpX bind symmetric heptameric rings of ClpP and interact with protein substrates. Subunits in the ClpX hexamer assume a spiral conformation and interact with two-residue segments of substrate in the axial channel, as observed for other AAA+ proteases and protein-remodeling machines. Strictly sequential models of ATP hydrolysis and a power stroke that moves two residues of the substrate per translocation step have been inferred from these structural features for other AAA+ unfoldases, but biochemical and single-molecule biophysical studies indicate that ClpXP operates by a probabilistic mechanism in which five to eight residues are translocated for each ATP hydrolyzed. We propose structure-based models that could account for the functional results.
- Department of Biology, Massachusetts Institute of Technology, Cambridge, United States.
Organizational Affiliation: