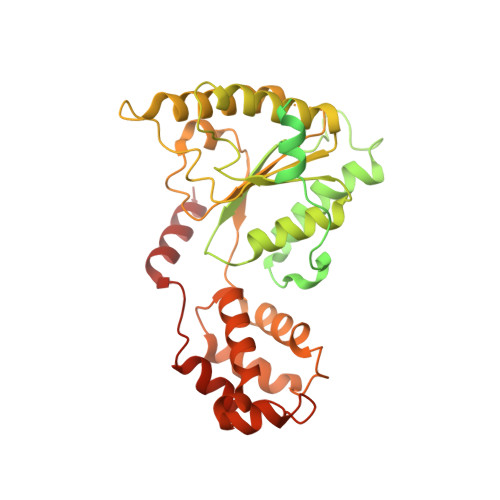
Structure of spastin bound to a glutamate-rich peptide implies a hand-over-hand mechanism of substrate translocation.
Han, H., Schubert, H.L., McCullough, J., Monroe, N., Purdy, M.D., Yeager, M., Sundquist, W.I., Hill, C.P.(2020) J Biological Chem 295: 435-443
- PubMed: 31767681 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.AC119.009890
- Primary Citation Related Structures:
6PEK, 6PEN - PubMed Abstract:
Many members of the AAA+ ATPase family function as hexamers that unfold their protein substrates. These AAA unfoldases include spastin, which plays a critical role in the architecture of eukaryotic cells by driving the remodeling and severing of microtubules, which are cytoskeletal polymers of tubulin subunits. Here, we demonstrate that a human spastin binds weakly to unmodified peptides from the C-terminal segment of human tubulin α1A/B. A peptide comprising alternating glutamate and tyrosine residues binds more tightly, which is consistent with the known importance of glutamylation for spastin microtubule severing activity. A cryo-EM structure of the spastin-peptide complex at 4.2 Å resolution revealed an asymmetric hexamer in which five spastin subunits adopt a helical, spiral staircase configuration that binds the peptide within the central pore, whereas the sixth subunit of the hexamer is displaced from the peptide/substrate, as if transitioning from one end of the helix to the other. This configuration differs from a recently published structure of spastin from Drosophila melanogaster , which forms a six-subunit spiral without a transitioning subunit. Our structure resembles other recently reported AAA unfoldases, including the meiotic clade relative Vps4, and supports a model in which spastin utilizes a hand-over-hand mechanism of tubulin translocation and microtubule remodeling.
- Department of Biochemistry, University of Utah School of Medicine, Salt Lake City, Utah 84112.
Organizational Affiliation: