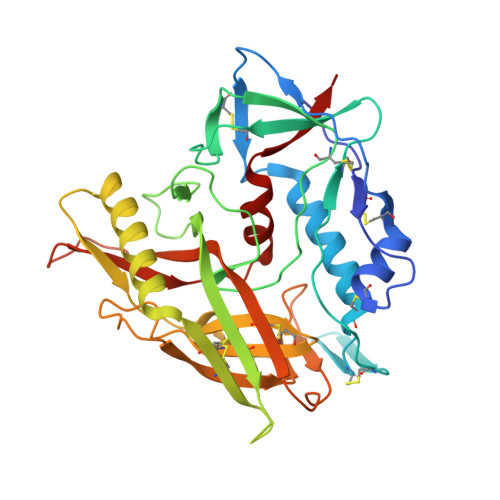
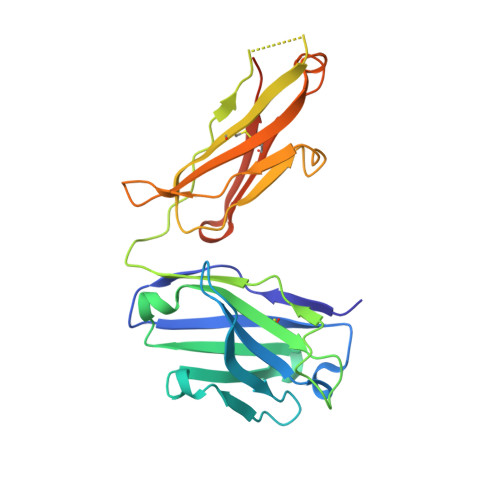
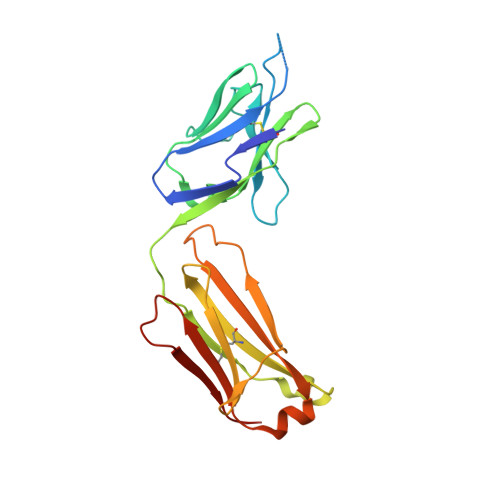
Overcoming Steric Restrictions of VRC01 HIV-1 Neutralizing Antibodies through Immunization.
Parks, K.R., MacCamy, A.J., Trichka, J., Gray, M., Weidle, C., Borst, A.J., Khechaduri, A., Takushi, B., Agrawal, P., Guenaga, J., Wyatt, R.T., Coler, R., Seaman, M., LaBranche, C., Montefiori, D.C., Veesler, D., Pancera, M., McGuire, A., Stamatatos, L.(2019) Cell Rep 29: 3060-3072.e7
- PubMed: 31801073 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.celrep.2019.10.071
- Primary Citation Related Structures:
6P8M, 6P8N - PubMed Abstract:
Broadly HIV-1 neutralizing VRC01 class antibodies target the CD4-binding site of Env. They are derived from VH1-2 ∗ 02 antibody heavy chains paired with rare light chains expressing 5-amino acid-long CDRL3s. They have been isolated from infected subjects but have not yet been elicited by immunization. Env-derived immunogens capable of binding the germline forms of VRC01 B cell receptors on naive B cells have been designed and evaluated in knockin mice. However, the elicited antibodies cannot bypass glycans present on the conserved position N276 of Env, which restricts access to the CD4-binding site. Efforts to guide the appropriate maturation of these antibodies by sequential immunization have not yet been successful. Here, we report on a two-step immunization scheme that leads to the maturation of VRC01-like antibodies capable of accommodating the N276 glycan and displaying autologous tier 2 neutralizing activities. Our results are relevant to clinical trials aiming to elicit VRC01 antibodies.
- Vaccines and Infectious Diseases Division, Fred Hutchinson Cancer Research Center, Seattle, WA, USA; Department of Global Health, University of Washington, Seattle, WA, USA.
Organizational Affiliation: