Structure of DNA polymerase III, beta subunit/ beta sliding clamp from Klebsiella pneumoniae, expressed with an N-terminal His-Smt3 fusion tag, in complex with Griselimycin
Abendroth, J., Lorimer, D.D., Horanyi, P.S., Edwards, T.E.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Ubiquitin-like protein SMT3,Beta sliding clamp | 462 | Saccharomyces cerevisiae S288C, Klebsiella pneumoniae IS22 This entity is chimeric | Mutation(s): 0 Gene Names: SMT3, YDR510W, D9719.15 | 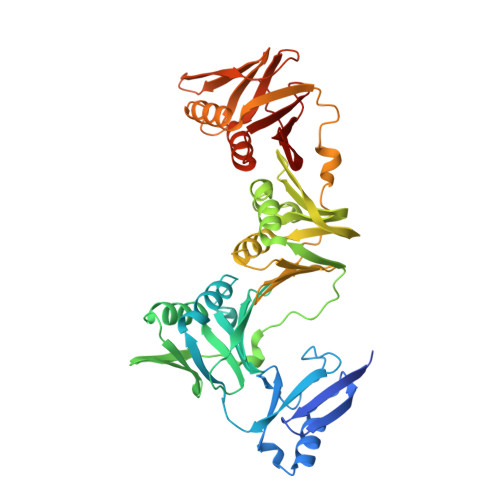 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q12306 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Griselimycin | 11 | Streptomyces muensis | Mutation(s): 0 | 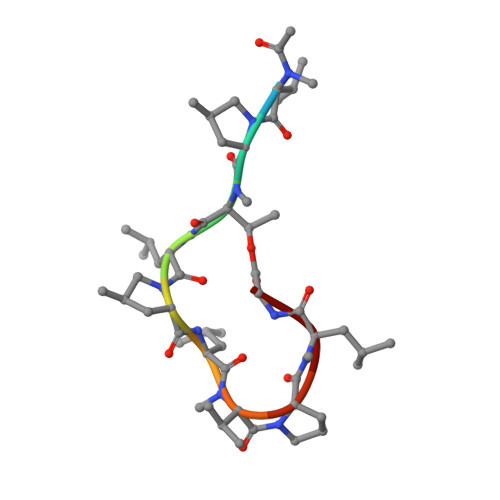 | |
| Ligands 4 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| PG5 Download:Ideal Coordinates CCD File | D [auth A] | 1-METHOXY-2-[2-(2-METHOXY-ETHOXY]-ETHANE C8 H18 O4 YFNKIDBQEZZDLK-UHFFFAOYSA-N |  | ||
| ACT Download:Ideal Coordinates CCD File | E [auth A] | ACETATE ION C2 H3 O2 QTBSBXVTEAMEQO-UHFFFAOYSA-M |  | ||
| CA Download:Ideal Coordinates CCD File | C [auth A] | CALCIUM ION Ca BHPQYMZQTOCNFJ-UHFFFAOYSA-N |  | ||
| CL Download:Ideal Coordinates CCD File | F [auth A] | CHLORIDE ION Cl VEXZGXHMUGYJMC-UHFFFAOYSA-M |  | ||
| Modified Residues 3 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| MLU Query on MLU | B | D-PEPTIDE LINKING | C7 H15 N O2 |  | -- |
| MP8 Query on MP8 | B | L-PEPTIDE LINKING | C6 H11 N O2 |  | PRO |
| MVA Query on MVA | B | L-PEPTIDE LINKING | C6 H13 N O2 |  | VAL |
| NZC Query on NZC | B | L-PEPTIDE LINKING | C5 H11 N O3 |  | THR |
| Entity ID: 2 | |||||
|---|---|---|---|---|---|
| ID | Chains | Name | Type/Class | 2D Diagram | 3D Interactions |
| PRD_002311 Query on PRD_002311 | B | ACE-MVA-MP8-NZC-LEU-MP8-LEU-MVA-PRO-MLU-GLY | Peptide-like / Inhibitor |  | |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 71.02 | α = 90 |
| b = 83.1 | β = 90 |
| c = 188.18 | γ = 90 |
| Software Name | Purpose |
|---|---|
| XDS | data reduction |
| XSCALE | data scaling |
| PHENIX | refinement |
| PDB_EXTRACT | data extraction |
| MoRDa | phasing |
| PARROT | phasing |
| ARP/wARP | model building |
| Coot | model building |