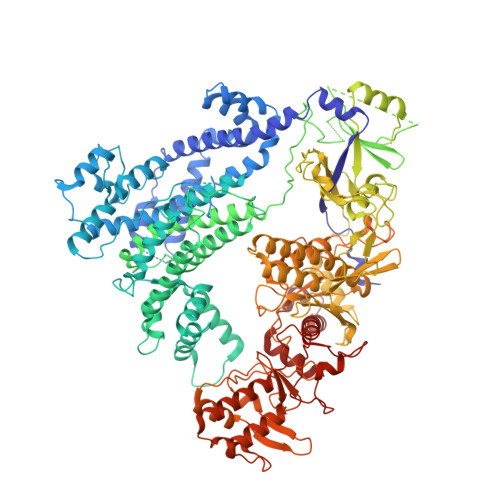
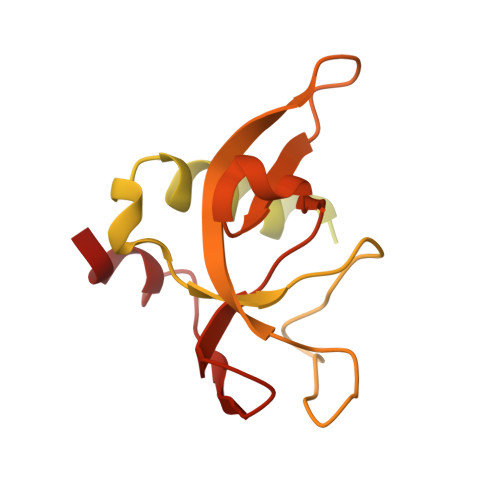
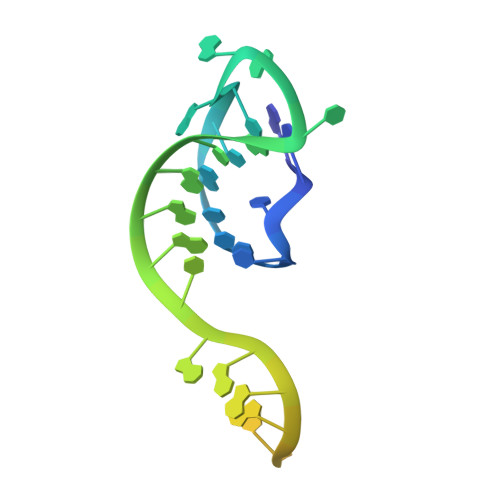
Structural basis for AcrVA4 inhibition of specific CRISPR-Cas12a.
Knott, G.J., Cress, B.F., Liu, J.J., Thornton, B.W., Lew, R.J., Al-Shayeb, B., Rosenberg, D.J., Hammel, M., Adler, B.A., Lobba, M.J., Xu, M., Arkin, A.P., Fellmann, C., Doudna, J.A.(2019) Elife 8
- PubMed: 31397669 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.49110
- Primary Citation Related Structures:
6P7M, 6P7N - PubMed Abstract:
CRISPR-Cas systems provide bacteria and archaea with programmable immunity against mobile genetic elements. Evolutionary pressure by CRISPR-Cas has driven bacteriophage to evolve small protein inhibitors, anti-CRISPRs (Acrs), that block Cas enzyme function by wide-ranging mechanisms. We show here that the inhibitor AcrVA4 uses a previously undescribed strategy to recognize the L. bacterium Cas12a (LbCas12a) pre-crRNA processing nuclease, forming a Cas12a dimer, and allosterically inhibiting DNA binding. The Ac. species Cas12a (AsCas12a) enzyme, widely used for genome editing applications, contains an ancestral helical bundle that blocks AcrVA4 binding and allows it to escape anti-CRISPR recognition. Using biochemical, microbiological, and human cell editing experiments, we show that Cas12a orthologs can be rendered either sensitive or resistant to AcrVA4 through rational structural engineering informed by evolution. Together, these findings explain a new mode of CRISPR-Cas inhibition and illustrate how structural variability in Cas effectors can drive opportunistic co-evolution of inhibitors by bacteriophage.
- Department of Molecular and Cell Biology, University of California, Berkeley, Berkeley, United States.
Organizational Affiliation: