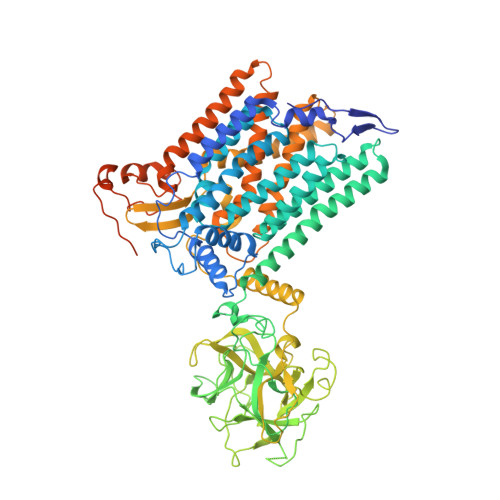
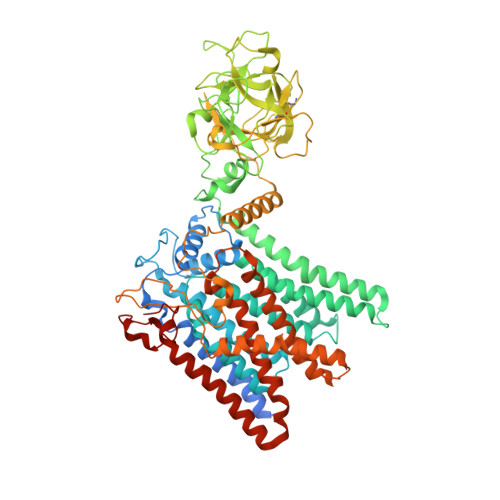
Structure of the eukaryotic protein O-mannosyltransferase Pmt1-Pmt2 complex.
Bai, L., Kovach, A., You, Q., Kenny, A., Li, H.(2019) Nat Struct Mol Biol 26: 704-711
- PubMed: 31285605 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-019-0262-6
- Primary Citation Related Structures:
6P25, 6P28, 6P2R - PubMed Abstract:
In eukaryotes, a nascent peptide entering the endoplasmic reticulum (ER) is scanned by two Sec61 translocon-associated large membrane machines for protein N-glycosylation and protein O-mannosylation, respectively. While the structure of the eight-protein oligosaccharyltransferase complex has been determined recently, the structures of mannosyltransferases of the PMT family, which are an integral part of ER protein homeostasis, are still unknown. Here we report cryo-EM structures of the Saccharomyces cerevisiae Pmt1-Pmt2 complex bound to a donor and an acceptor peptide at 3.2-Å resolution, showing that each subunit contains 11 transmembrane helices and a lumenal β-trefoil fold termed the MIR domain. The structures reveal the substrate recognition model and confirm an inverting mannosyl-transferring reaction mechanism by the enzyme complex. Furthermore, we found that the transmembrane domains of Pmt1 and Pmt2 share a structural fold with the catalytic subunits of oligosaccharyltransferases, confirming a previously proposed evolutionary relationship between protein O-mannosylation and protein N-glycosylation.
- Structural Biology Program, Van Andel Research Institute, Grand Rapids, MI, USA.
Organizational Affiliation: