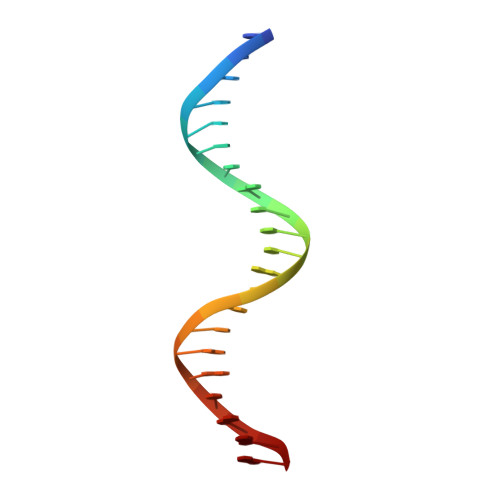
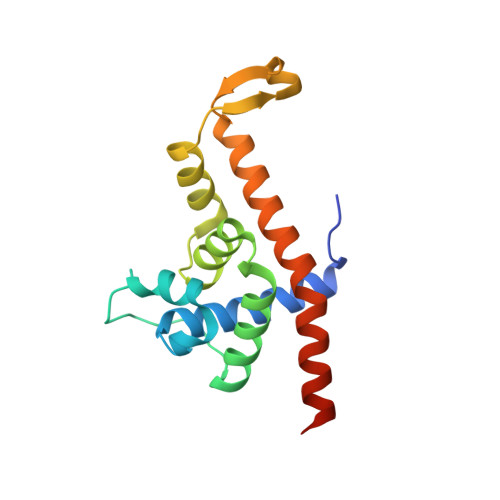
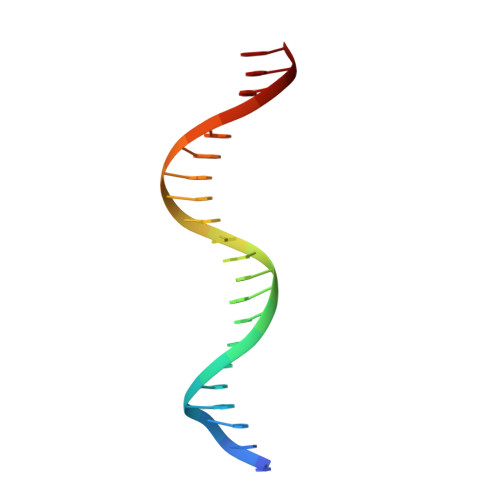
Structural basis of Q-dependent antitermination.
Yin, Z., Kaelber, J.T., Ebright, R.H.(2019) Proc Natl Acad Sci U S A 116: 18384-18390
- PubMed: 31455742 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1909801116
- Primary Citation Related Structures:
6P18, 6P19, 6P1A, 6P1B, 6P1C - PubMed Abstract:
Lambdoid bacteriophage Q protein mediates the switch from middle to late bacteriophage gene expression by enabling RNA polymerase (RNAP) to read through transcription terminators preceding bacteriophage late genes. Q loads onto RNAP engaged in promoter-proximal pausing at a Q binding element (QBE) and adjacent sigma-dependent pause element (SDPE) to yield a Q-loading complex, and Q subsequently translocates with RNAP as a pausing-deficient, termination-deficient Q-loaded complex. Here, we report high-resolution structures of 4 states on the pathway of antitermination by Q from bacteriophage 21 (Q21): Q21, the Q21-QBE complex, the Q21-loading complex, and the Q21-loaded complex. The results show that Q21 forms a torus, a "nozzle," that narrows and extends the RNAP RNA-exit channel, extruding topologically linked single-stranded RNA and preventing the formation of pause and terminator hairpins.
- Waksman Institute, Rutgers University, Piscataway, NJ 08854.
Organizational Affiliation: