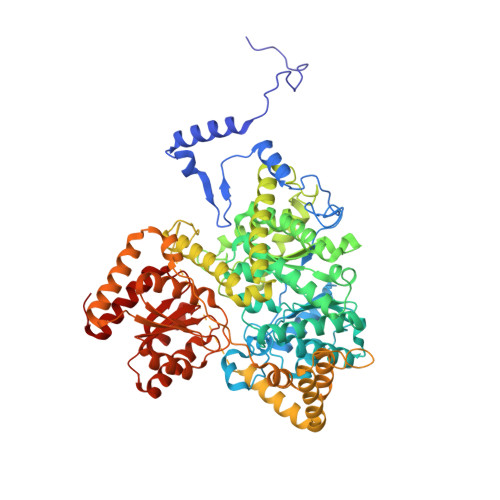
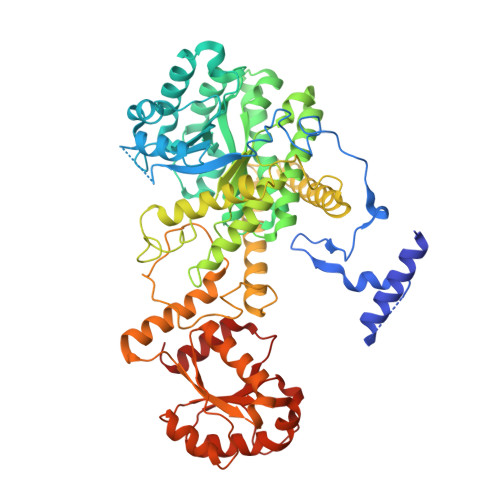
Itaconyl-CoA forms a stable biradical in methylmalonyl-CoA mutase and derails its activity and repair.
Ruetz, M., Campanello, G.C., Purchal, M., Shen, H., McDevitt, L., Gouda, H., Wakabayashi, S., Zhu, J., Rubin, E.J., Warncke, K., Mootha, V.K., Koutmos, M., Banerjee, R.(2019) Science 366: 589-593
- PubMed: 31672889 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/science.aay0934
- Primary Citation Related Structures:
6OXC, 6OXD - PubMed Abstract:
Itaconate is an immunometabolite with both anti-inflammatory and bactericidal effects. Its coenzyme A (CoA) derivative, itaconyl-CoA, inhibits B 12 -dependent methylmalonyl-CoA mutase (MCM) by an unknown mechanism. We demonstrate that itaconyl-CoA is a suicide inactivator of human and Mycobacterium tuberculosis MCM, which forms a markedly air-stable biradical adduct with the 5'-deoxyadenosyl moiety of the B 12 coenzyme. Termination of the catalytic cycle in this way impairs communication between MCM and its auxiliary repair proteins. Crystallography and spectroscopy of the inhibited enzyme are consistent with a metal-centered cobalt radical ~6 angstroms away from the tertiary carbon-centered radical and suggest a means of controlling radical trajectories during MCM catalysis. Mycobacterial MCM thus joins enzymes in the glyoxylate shunt and the methylcitrate cycle as targets of itaconate in pathogen propionate metabolism.
- Department of Biological Chemistry, University of Michigan, Ann Arbor, MI 48109, USA.
Organizational Affiliation: