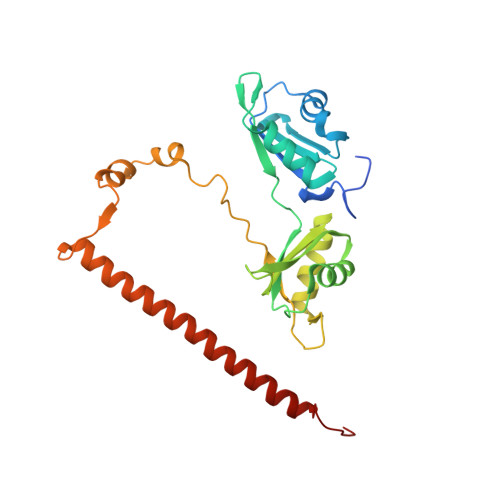
Structural basis of the zinc-induced cytoplasmic aggregation of the RNA-binding protein SFPQ.
Huang, J., Ringuet, M., Whitten, A.E., Caria, S., Lim, Y.W., Badhan, R., Anggono, V., Lee, M.(2020) Nucleic Acids Res 48: 3356-3365
- PubMed: 32034402 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkaa076
- Primary Citation Related Structures:
6OWJ - PubMed Abstract:
SFPQ is a ubiquitous nuclear RNA-binding protein implicated in many aspects of RNA biogenesis. Importantly, nuclear depletion and cytoplasmic accumulation of SFPQ has been linked to neuropathological conditions such as Alzheimer's disease (AD) and amyotrophic lateral sclerosis (ALS). Here, we describe a molecular mechanism by which SFPQ is mislocalized to the cytoplasm. We report an unexpected discovery of the infinite polymerization of SFPQ that is induced by zinc binding to the protein. The crystal structure of human SFPQ in complex with zinc at 1.94 Å resolution reveals intermolecular interactions between SFPQ molecules that are mediated by zinc. As anticipated from the crystal structure, the application of zinc to primary cortical neurons induced the cytoplasmic accumulation and aggregation of SFPQ. Mutagenesis of the three zinc-coordinating histidine residues resulted in a significant reduction in the zinc-binding affinity of SFPQ in solution and the zinc-induced cytoplasmic aggregation of SFPQ in cultured neurons. Taken together, we propose that dysregulation of zinc availability and/or localization in neuronal cells may represent a mechanism for the imbalance in the nucleocytoplasmic distribution of SFPQ, which is an emerging hallmark of neurodegenerative diseases including AD and ALS.
- Department of Biochemistry and Genetics, La Trobe Institute for Molecular Science, La Trobe University, Melbourne, Victoria 3086, Australia.
Organizational Affiliation: