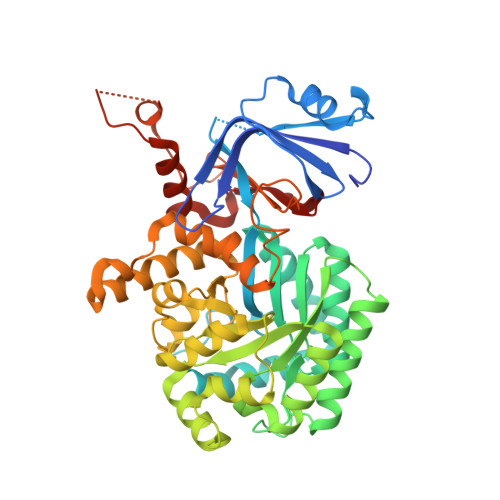
Structural Determinants for Substrate Selectivity in Guanine Deaminase Enzymes of the Amidohydrolase Superfamily.
Shek, R., Hilaire, T., Sim, J., French, J.B.(2019) Biochemistry 58: 3280-3292
- PubMed: 31283204 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.biochem.9b00341
- Primary Citation Related Structures:
6OH9, 6OHA, 6OHB, 6OHC - PubMed Abstract:
Guanine deaminase is a metabolic enzyme, found in all forms of life, which catalyzes the conversion of guanine to xanthine. Despite the availability of several crystal structures, the molecular determinants of substrate orientation and mechanism remain to be elucidated for the amidohydrolase family of guanine deaminase enzymes. Here, we report the crystal structures of Escherichia coli and Saccharomyces cerevisiae guanine deaminase enzymes (EcGuaD and Gud1, respectively), both members of the amidohydrolase superfamily. EcGuaD and Gud1 retain the overall TIM barrel tertiary structure conserved among amidohydrolase enzymes. Both proteins also possess a single zinc cation with trigonal bipyrimidal coordination geometry within their active sites. We also determined a liganded structure of Gud1 bound to the product, xanthine. Analysis of this structure, along with kinetic data of native and site-directed mutants of EcGuaD, identifies several key residues that are responsible for substrate recognition and catalysis. In addition, after a small library of compounds had been screened, two guanine derivatives, 8-azaguanine and 1-methylguanine, were identified as EcGuaD substrates. Interestingly, both EcGuaD and Gud1 also exhibit secondary ammeline deaminase activity. Overall, this work details key structural features of substrate recognition and catalysis of the amidohydrolase family of guanine deaminase enzymes in support of our long-term goal to engineer these enzymes with altered activity and substrate specificity.
- Department of Biochemistry and Cell Biology , Stony Brook University , Stony Brook , New York 11794 , United States.
Organizational Affiliation: