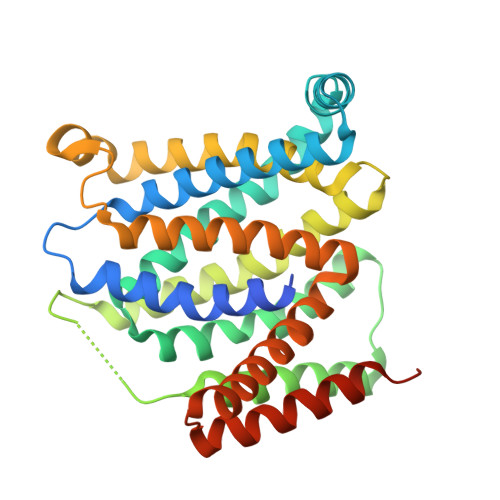
Structural basis for mammalian nucleotide sugar transport.
Ahuja, S., Whorton, M.R.(2019) Elife 8
- PubMed: 30985278 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.45221
- Primary Citation Related Structures:
6OH2, 6OH3, 6OH4 - PubMed Abstract:
Nucleotide-sugar transporters (NSTs) are critical components of the cellular glycosylation machinery. They transport nucleotide-sugar conjugates into the Golgi lumen, where they are used for the glycosylation of proteins and lipids, and they then subsequently transport the nucleotide monophosphate byproduct back to the cytoplasm. Dysregulation of human NSTs causes several debilitating diseases, and NSTs are virulence factors for many pathogens. Here we present the first crystal structures of a mammalian NST, the mouse CMP-sialic acid transporter (mCST), in complex with its physiological substrates CMP and CMP-sialic acid. Detailed visualization of extensive protein-substrate interactions explains the mechanisms governing substrate selectivity. Further structural analysis of mCST's unique lumen-facing partially-occluded conformation, coupled with the characterization of substrate-induced quenching of mCST's intrinsic tryptophan fluorescence, reveals the concerted conformational transitions that occur during substrate transport. These results provide a framework for understanding the effects of disease-causing mutations and the mechanisms of this diverse family of transporters.
- Vollum Institute, Oregon Health & Science University, Portland, United States.
Organizational Affiliation: