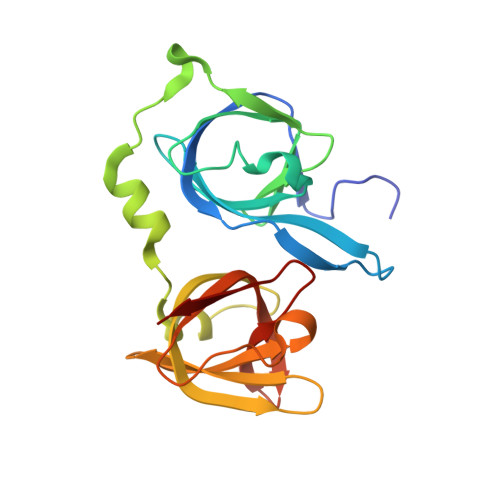
Structural analysis of the HIN1 domain of interferon-inducible protein 204.
Tian, Y., Yin, Q.(2019) Acta Crystallogr F Struct Biol Commun 75: 455-460
- PubMed: 31204693 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X19007167
- Primary Citation Related Structures:
6OE9 - PubMed Abstract:
Interferon-inducible protein 204 (p204) binds to microbial DNA to elicit inflammatory responses and induce interferon production. p204 also modulates cell proliferation and differentiation by regulating various transcription factors. The C-terminal HIN domains in p204 are believed to be responsible for DNA binding, but the binding mode is not fully understood. The DNA-binding affinity of the p204 HIN1 domain has been characterized and its crystal structure has been determined, providing insight into its interaction with DNA. Surface-charge distribution together with sequence alignment suggests that the p204 HIN domain uses its L12 and L45 loops for DNA binding.
- Department of Biological Science, Florida State University, Tallahassee, FL 32306, USA.
Organizational Affiliation: