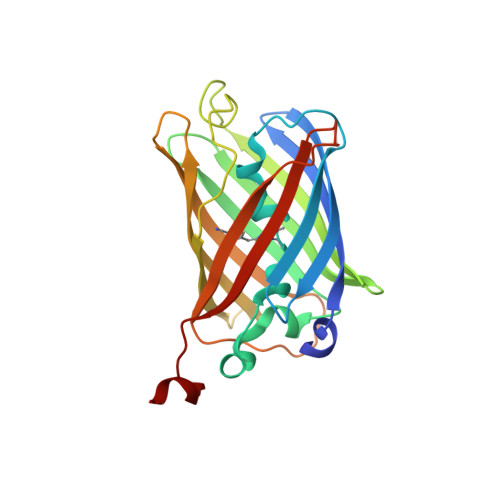
Structural and spectrophotometric investigation of two unnatural amino-acid altered chromophores in the superfolder green fluorescent protein
Olenginski, G.M., Piacentini, J., Harris, D.R., Runko, N., Papoutsis, B.M., Alter, J.R., Hess, K.R., Brewer, S.H., Phillips-Piro, C.M.(2021) Acta Crystallogr D Biol Crystallogr