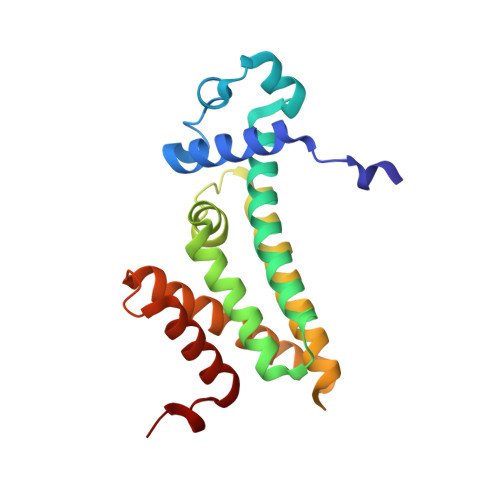
Mycobacterium tuberculosis FasR senses long fatty acyl-CoA through a tunnel and a hydrophobic transmission spine.
Lara, J., Diacovich, L., Trajtenberg, F., Larrieux, N., Malchiodi, E.L., Fernandez, M.M., Gago, G., Gramajo, H., Buschiazzo, A.(2020) Nat Commun 11: 3703-3703
- PubMed: 32710080 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-020-17504-x
- Primary Citation Related Structures:
6O6N, 6O6O, 6O6P - PubMed Abstract:
Mycobacterium tuberculosis is a pathogen with a unique cell envelope including very long fatty acids, implicated in bacterial resistance and host immune modulation. FasR is a TetR-like transcriptional activator that plays a central role in sensing mycobacterial long-chain fatty acids and regulating lipid biosynthesis. Here we disclose crystal structures of M. tuberculosis FasR in complex with acyl effector ligands and with DNA, uncovering its molecular sensory and switching mechanisms. A long tunnel traverses the entire effector-binding domain, enabling long fatty acyl effectors to bind. Only when the tunnel is entirely occupied, the protein dimer adopts a rigid configuration with its DNA-binding domains in an open state, leading to DNA dissociation. The protein-folding hydrophobic core connects the two domains, and is completed into a continuous spine when the effector binds. Such a transmission spine is conserved in a large number of TetR-like regulators, offering insight into effector-triggered allosteric functional control.
- Laboratory of Physiology and Genetics of Actinomycetes, Instituto de Biología Molecular y Celular de Rosario (IBR-CONICET), Facultad de Ciencias Bioquímicas y Farmacéuticas, Universidad Nacional de Rosario, Rosario, 2000, Argentina.
Organizational Affiliation: