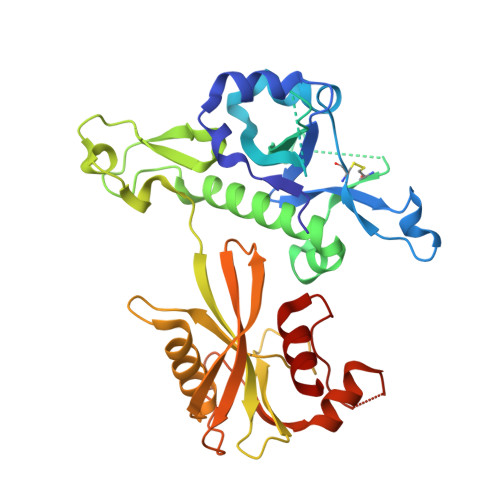
Structural and mechanistic basis of mammalian Nudt12 RNA deNADding.
Grudzien-Nogalska, E., Wu, Y., Jiao, X., Cui, H., Mateyak, M.K., Hart, R.P., Tong, L., Kiledjian, M.(2019) Nat Chem Biol 15: 575-582
- PubMed: 31101919 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41589-019-0293-7
- Primary Citation Related Structures:
6O3P - PubMed Abstract:
We recently demonstrated that mammalian cells harbor nicotinamide adenine dinucleotide (NAD)-capped messenger RNAs that are hydrolyzed by the DXO deNADding enzyme. Here, we report that the Nudix protein Nudt12 is a second mammalian deNADding enzyme structurally and mechanistically distinct from DXO and targeting different RNAs. The crystal structure of mouse Nudt12 in complex with the deNADding product AMP and three Mg 2+ ions at 1.6 Å resolution provides insights into the molecular basis of the deNADding activity in the NAD pyrophosphate. Disruption of the Nudt12 gene stabilizes transfected NAD-capped RNA in cells, and its endogenous NAD-capped mRNA targets are enriched in those encoding proteins involved in cellular energetics. Furthermore, exposure of cells to nutrient or environmental stress manifests changes in NAD-capped RNA levels that are selectively responsive to Nudt12 or DXO, respectively, indicating an association of deNADding to cellular metabolism.
- Department Cell Biology and Neuroscience, Rutgers University, Piscataway, NJ, USA.
Organizational Affiliation: